Main Spatial R Packages
New ecosystem
Old ecosystem
2026-03-25
What This Workshop Is
What This Workshop Is Not
By the end of the workshop, participants should be able to use R as a GIS, work operationally with common spatial data manipulations, and have the right tools in hand to keep learning.
Materials
Exercises
The workshop alternates between concise instructional material and applied exercises, so participants can move directly from concepts to practice.
| Object type | Description |
sf
|
stars
|
terra
|
|---|---|---|---|---|
| Vector | Where are the features? | |||
|
|
Discrete locations with coordinates and attributes. | ✓ | ✓ | |
|
|
Linear features such as roads, rivers, or tracks. | ✓ | ✓ | |
|
|
Areas such as lakes, study zones, or administrative boundaries. | ✓ | ✓ | |
| Raster | What is the value across space? | |||
|
|
Gridded surfaces with values stored in cells. | ✓ | ✓ |
New ecosystem
stars 🌐
sfterra 🌐
Old ecosystem
Data manipulation and measurements
tidyverse 🌐
tidyterra 🌐
terraterraunits 🌐
sfterra, returns numeric measurementsVisualization
ggplot2 🌐
mapview 🌐
tmap 🌐
These are examples of particularly relevant accompanying packages. The spatial ecosystem in R is broad and active, and is not limited to the packages shown here.
What are IDEs? Integrated Development Environments
RStudio 🌐
VS Code 🌐
Positron 🌐
The goal of this workshop is not to teach a specific IDE. Use the environment that lets you work comfortably with scripts, objects, plots, and spatial outputs.
What are pipes?
Pipes pass the output of one step into the next, making multi-step workflows easier to read and write. We mention them explicitly because they are used throughout the workshop.
Choose Your Tool(s)
Navigating through the code in the presentation
The slides often show the same operation twice: once with sf/stars, and once with terra.
You do not need to follow both versions at once. Use the tabs to stay in the ecosystem you chose for the workshop.
Choose Your Route
Do both levels!
More advanced participants can complete both workflows, as there will be advanced workflows available with both points and lines.
sf & terra
Simple feature collection with 1 feature and 1 field
Geometry type: POLYGON
Dimension: XY
Bounding box: xmin: -73 ymin: 46.55 xmax: -72.6 ymax: 47
Geodetic CRS: WGS 84
name geom
1 study_area POLYGON ((-73 46.55, -72.6 ...Simple feature collection with 4 features and 2 fields
Geometry type: POLYGON
Dimension: XY
Bounding box: xmin: -73 ymin: 46.55 xmax: -72.6 ymax: 47
Geodetic CRS: WGS 84
zone_id zone_type geom
1 NW forest POLYGON ((-73 46.775, -72.8...
2 NE urban POLYGON ((-72.8 46.775, -72...
3 SW wetland POLYGON ((-73 46.55, -72.8 ...
4 SE agriculture POLYGON ((-72.8 46.55, -72....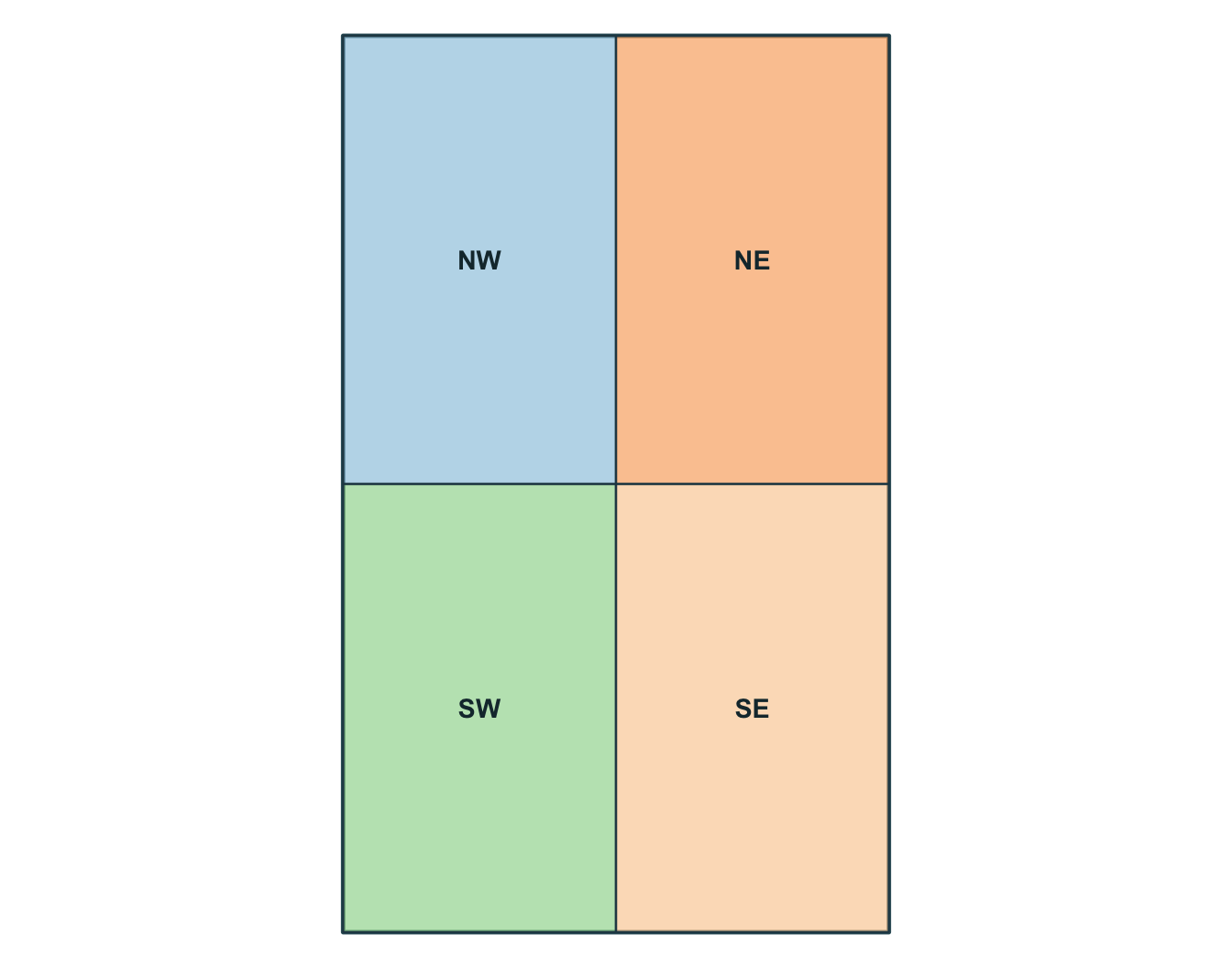
Simple feature collection with 6 features and 8 fields
Geometry type: POINT
Dimension: XY
Bounding box: xmin: -72.92 ymin: 46.7 xmax: -72.72 ymax: 46.94
Geodetic CRS: WGS 84
obs_id track_id seq timestamp lon lat category value
1 OBS01 A 1 2026-03-01 08:00:00 -72.92 46.78 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 -72.86 46.83 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 -72.79 46.88 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 -72.72 46.94 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 -72.88 46.70 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 -72.83 46.76 agriculture 13
geometry
1 POINT (-72.92 46.78)
2 POINT (-72.86 46.83)
3 POINT (-72.79 46.88)
4 POINT (-72.72 46.94)
5 POINT (-72.88 46.7)
6 POINT (-72.83 46.76)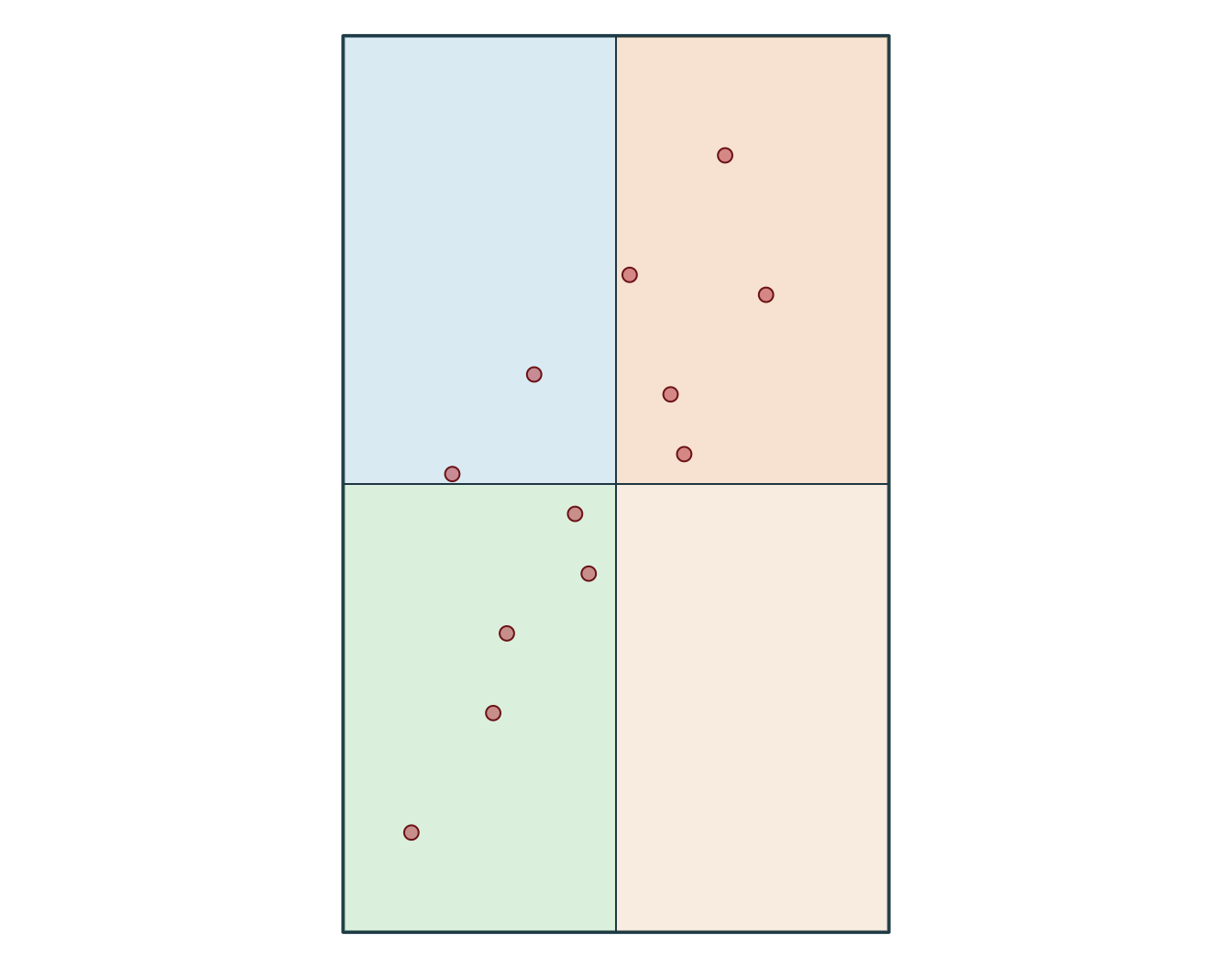
obs_id track_id seq timestamp category value
1 OBS01 A 1 2026-03-01 08:00:00 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 agriculture 13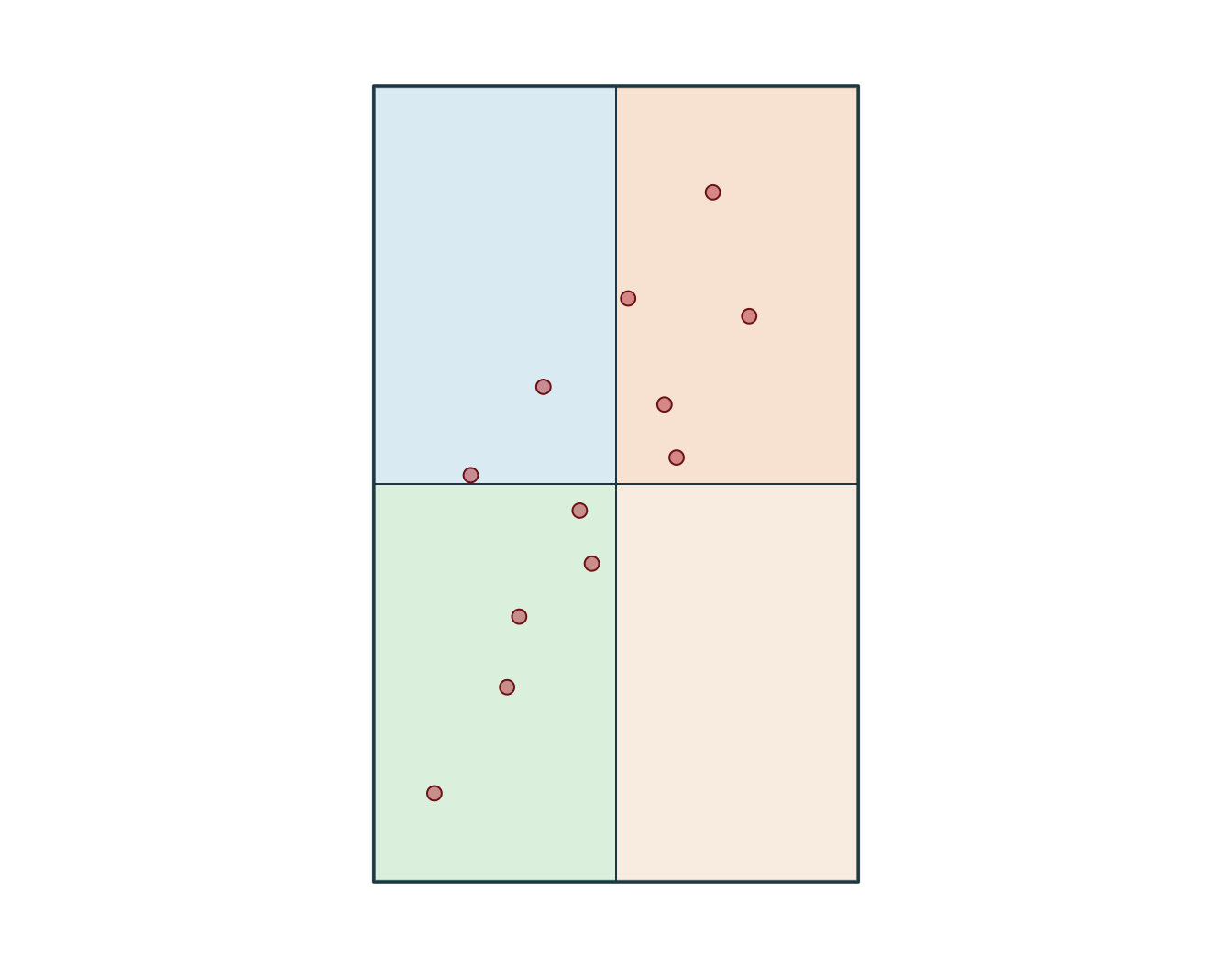
[1] POLYGON POLYGON POLYGON POLYGON
18 Levels: GEOMETRY POINT LINESTRING POLYGON MULTIPOINT ... TRIANGLE[1] "WGS 84" xmin ymin xmax ymax
-73.00 46.55 -72.60 47.00 Spatial projections
CRS information tells you how coordinates are referenced on the Earth. When you need to inspect or identify a projection, epsg.io is a useful lookup tool.
[1] "EPSG:32198" xmin ymin xmax ymax
-340347.4 299996.5 -318731.5 336605.7 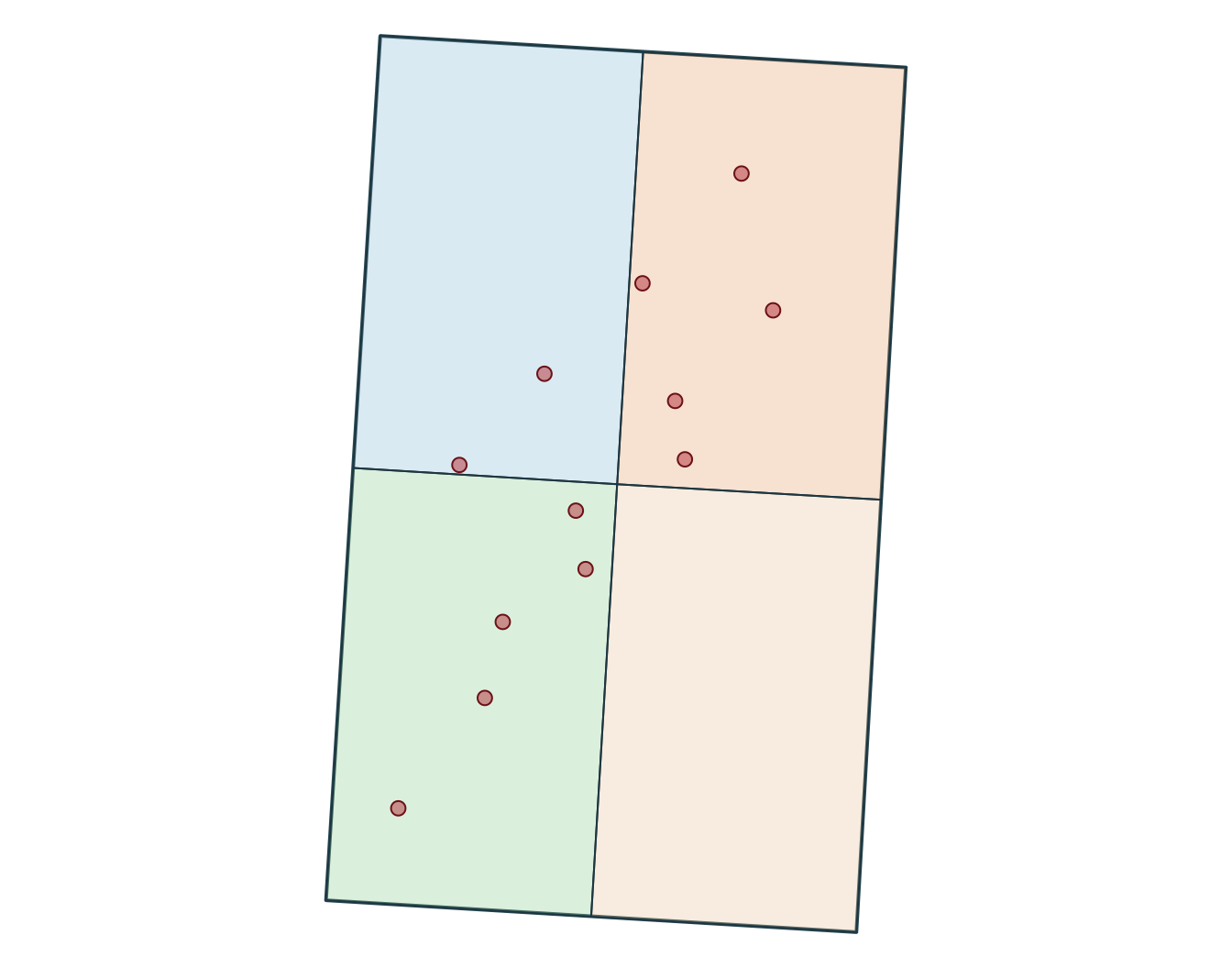
[1] "+proj=lcc +lat_0=44 +lon_0=-68.5 +lat_1=60 +lat_2=46 +x_0=0 +y_0=0 +datum=NAD83 +units=m +no_defs"SpatExtent : -340347.355525386, -318731.482880504, 299996.469057655, 336605.697992394 (xmin, xmax, ymin, ymax)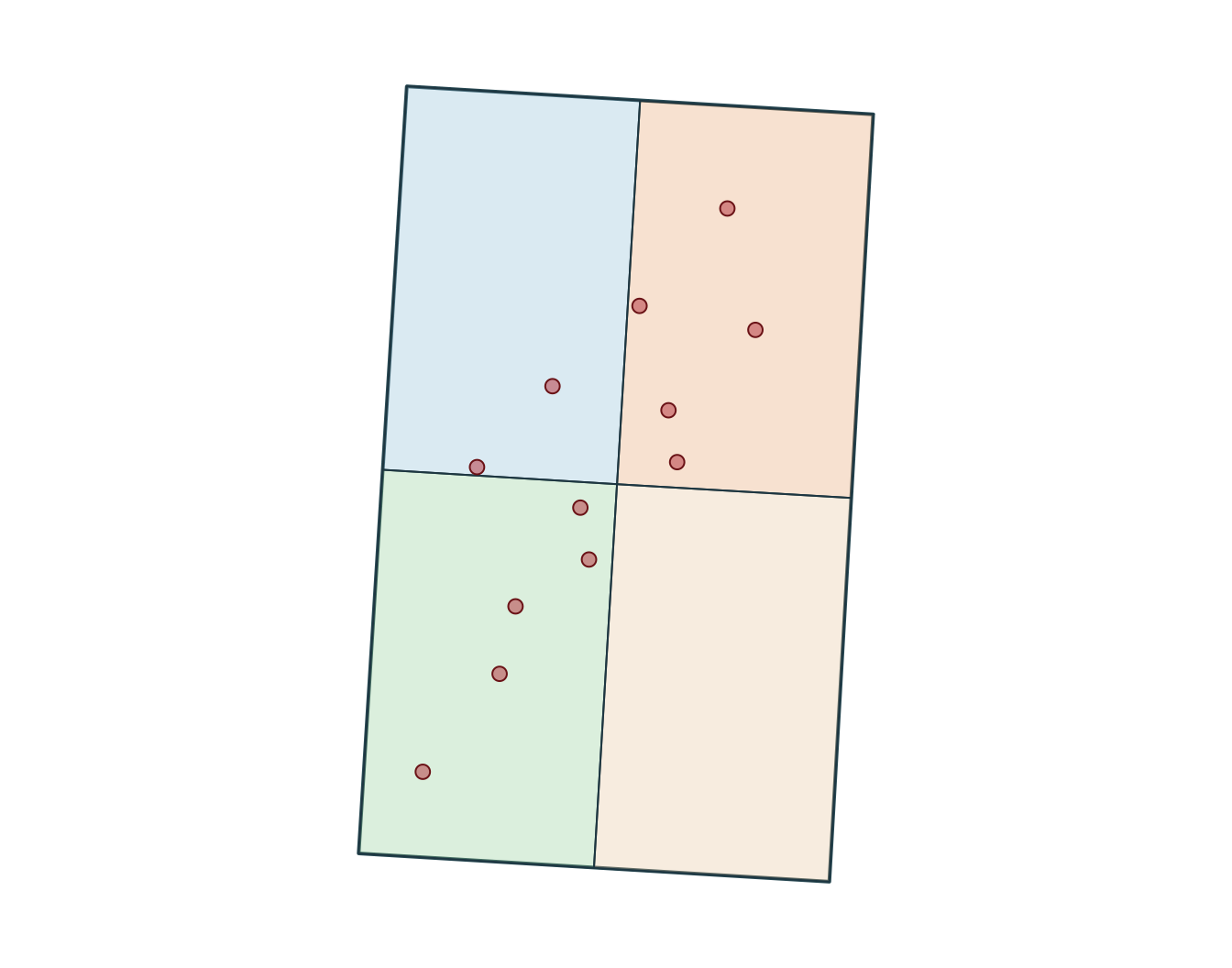
Measurements
Use a suitable projected CRS when you need interpretable distances, lengths, or areas.
Import, inspect, transform, export
Bonus: use R as a converter
Simple feature collection with 3 features and 1 field
Geometry type: LINESTRING
Dimension: XY
Bounding box: xmin: -72.95 ymin: 46.6 xmax: -72.69 ymax: 46.94
Geodetic CRS: WGS 84
# A tibble: 3 × 2
track_id geometry
<chr> <LINESTRING [°]>
1 A (-72.92 46.78, -72.86 46.83, -72.79 46.88, -72.72 46.94)
2 B (-72.88 46.7, -72.83 46.76, -72.76 46.82, -72.69 46.87)
3 C (-72.95 46.6, -72.89 46.66, -72.82 46.73, -72.75 46.79)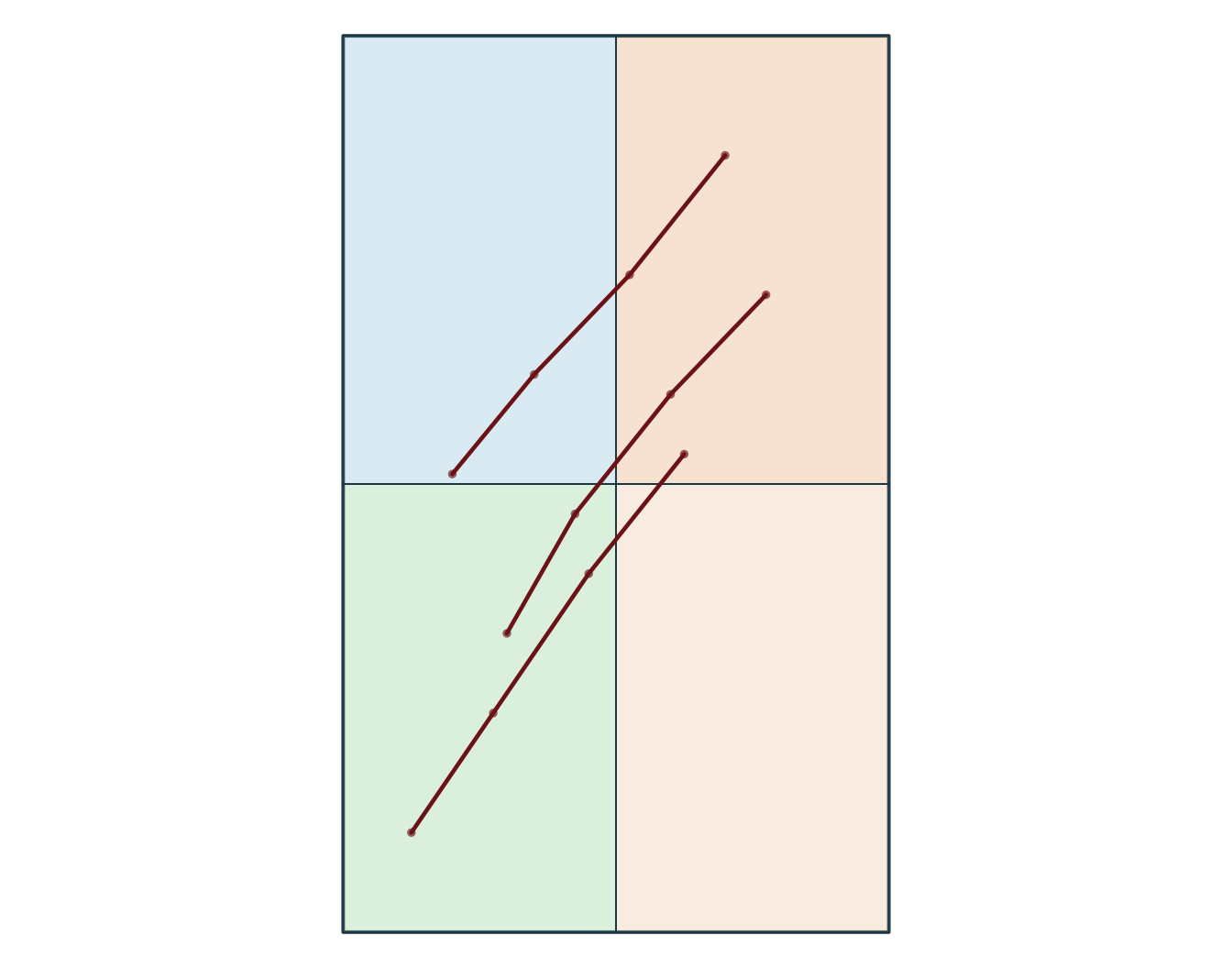
object file format
1 mapview sf map fileea35da38cec.html htmlShare your maps!
Interactive mapview outputs can be saved as standalone HTML files and shared like any other web document.
Import, inspect, transform, export, visualize
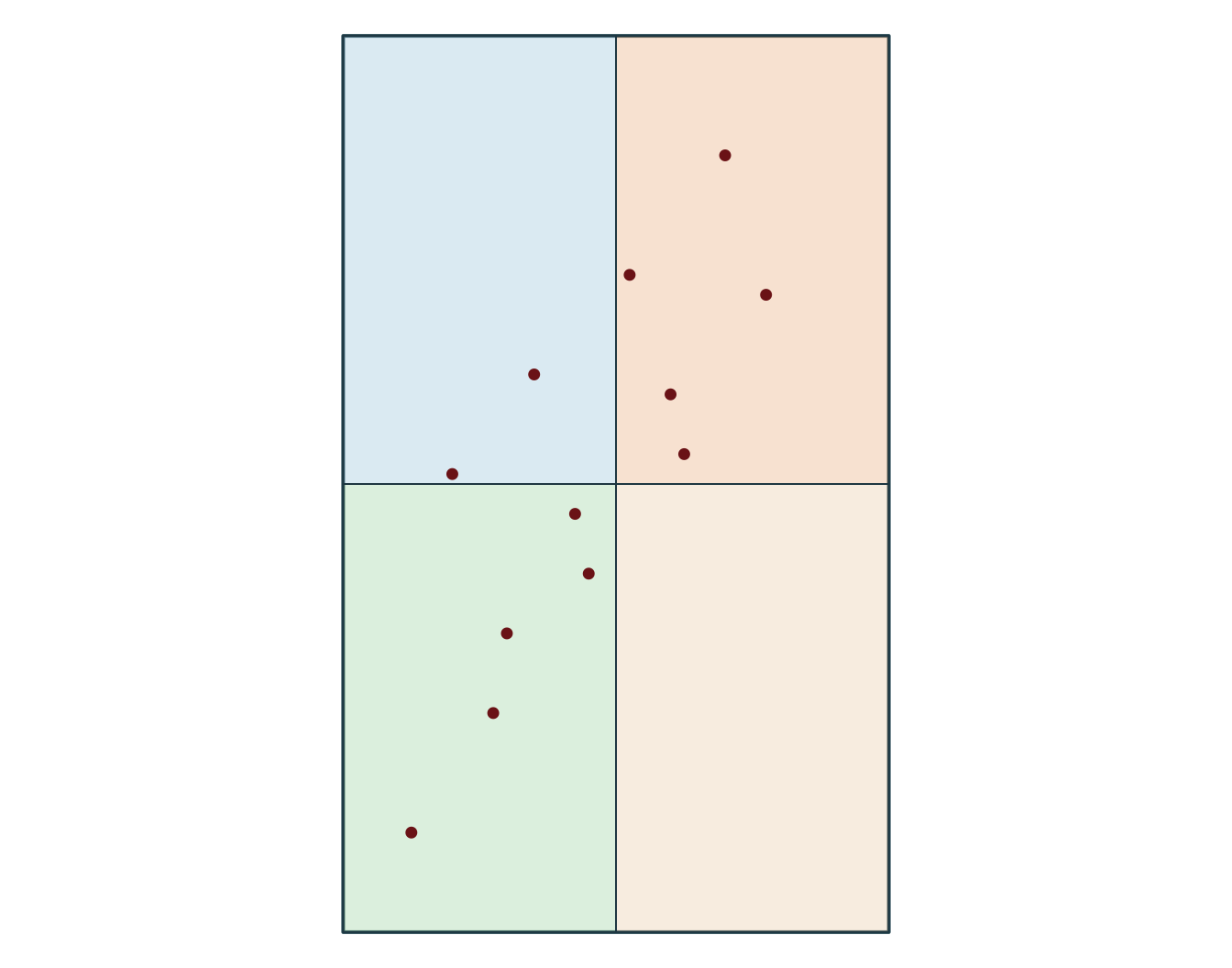
Simple feature collection with 5 features and 8 fields
Geometry type: POINT
Dimension: XY
Bounding box: xmin: -72.79 ymin: 46.79 xmax: -72.69 ymax: 46.94
Geodetic CRS: WGS 84
obs_id track_id seq timestamp lon lat category value
1 OBS03 A 3 2026-03-01 10:00:00 -72.79 46.88 forest 21
2 OBS04 A 4 2026-03-01 11:00:00 -72.72 46.94 forest 24
3 OBS07 B 3 2026-03-02 10:00:00 -72.76 46.82 wetland 17
4 OBS08 B 4 2026-03-02 11:00:00 -72.69 46.87 wetland 18
5 OBS12 C 4 2026-03-03 11:00:00 -72.75 46.79 urban 19
geometry
1 POINT (-72.79 46.88)
2 POINT (-72.72 46.94)
3 POINT (-72.76 46.82)
4 POINT (-72.69 46.87)
5 POINT (-72.75 46.79)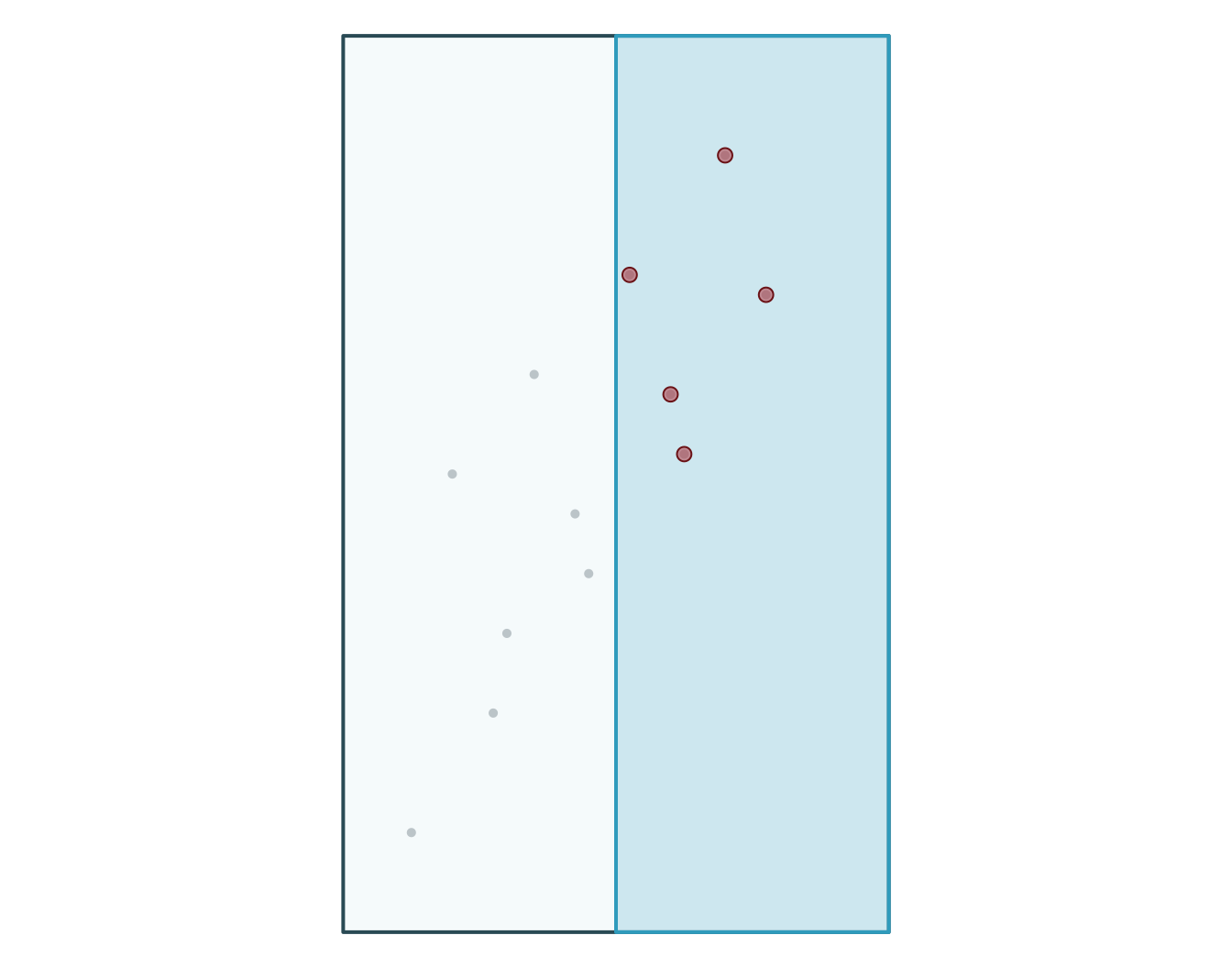
obs_id track_id seq timestamp category value
1 OBS03 A 3 2026-03-01 10:00:00 forest 21
2 OBS04 A 4 2026-03-01 11:00:00 forest 24
3 OBS07 B 3 2026-03-02 10:00:00 wetland 17
4 OBS08 B 4 2026-03-02 11:00:00 wetland 18
5 OBS12 C 4 2026-03-03 11:00:00 urban 19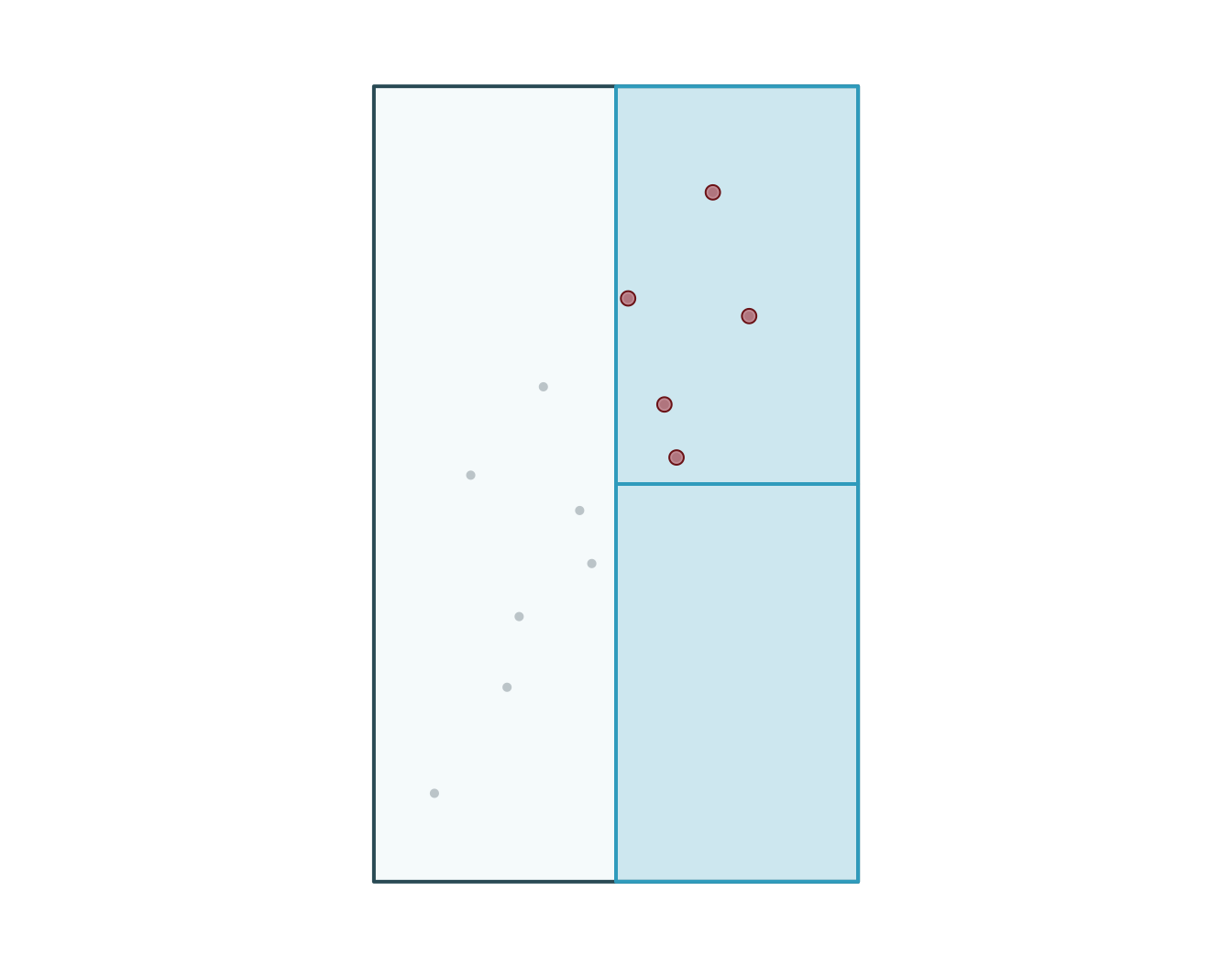
zone_id area_km2
1 NW 380.0
2 NE 380.0
3 SW 381.9
4 SE 381.9 track_id length_km
1 A 23.4
2 B 23.9
3 C 26.1 obs_id boundary_km
1 OBS01 6.1
2 OBS02 10.7
3 OBS03 13.3
4 OBS04 6.7
5 OBS05 9.2
6 OBS06 13.0 zone_id area_km2
1 NW 381.3
2 NE 381.3
3 SW 382.9
4 SE 382.9 track_id length_km
1 A 23.4
2 B 23.9
3 C 26.1 obs_id boundary_km
1 OBS01 6.1
2 OBS02 10.7
3 OBS03 13.3
4 OBS04 6.7
5 OBS05 9.2
6 OBS06 13.0Note
sf often returns unit-aware measurements. terra often returns numeric values whose meaning depends on the CRS and function used.
obs_id track_id seq timestamp lon lat category value
1 OBS01 A 1 2026-03-01 08:00:00 -72.92 46.78 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 -72.86 46.83 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 -72.79 46.88 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 -72.72 46.94 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 -72.88 46.70 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 -72.83 46.76 agriculture 13
zone_id zone_type
1 NW forest
2 NW forest
3 NE urban
4 NE urban
5 SW wetland
6 SW wetland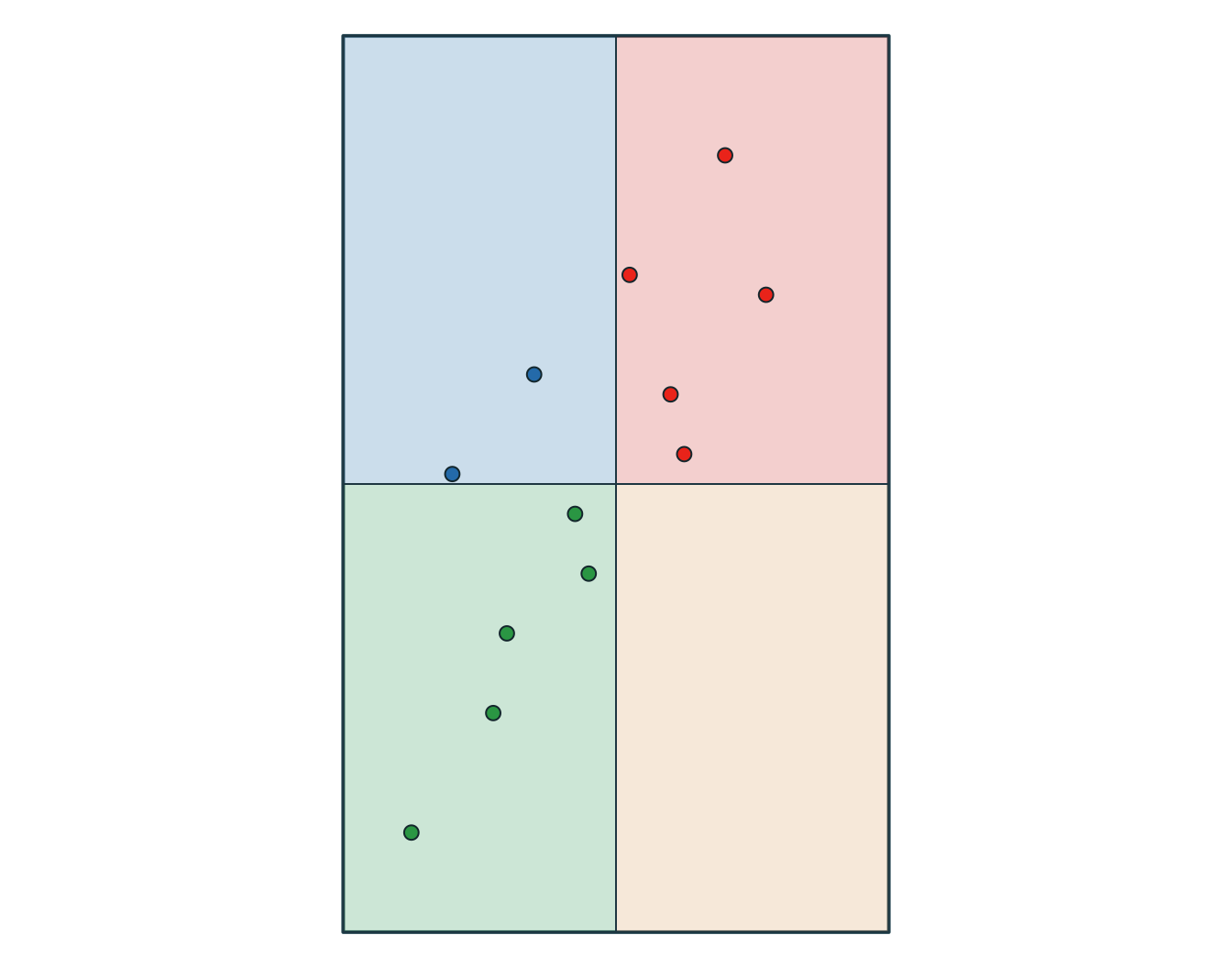
obs_id track_id seq timestamp category value zone_id zone_type
1 OBS01 A 1 2026-03-01 08:00:00 urban 14 NW forest
2 OBS02 A 2 2026-03-01 09:00:00 urban 16 NW forest
3 OBS03 A 3 2026-03-01 10:00:00 forest 21 NE urban
4 OBS04 A 4 2026-03-01 11:00:00 forest 24 NE urban
5 OBS05 B 1 2026-03-02 08:00:00 agriculture 11 SW wetland
6 OBS06 B 2 2026-03-02 09:00:00 agriculture 13 SW wetland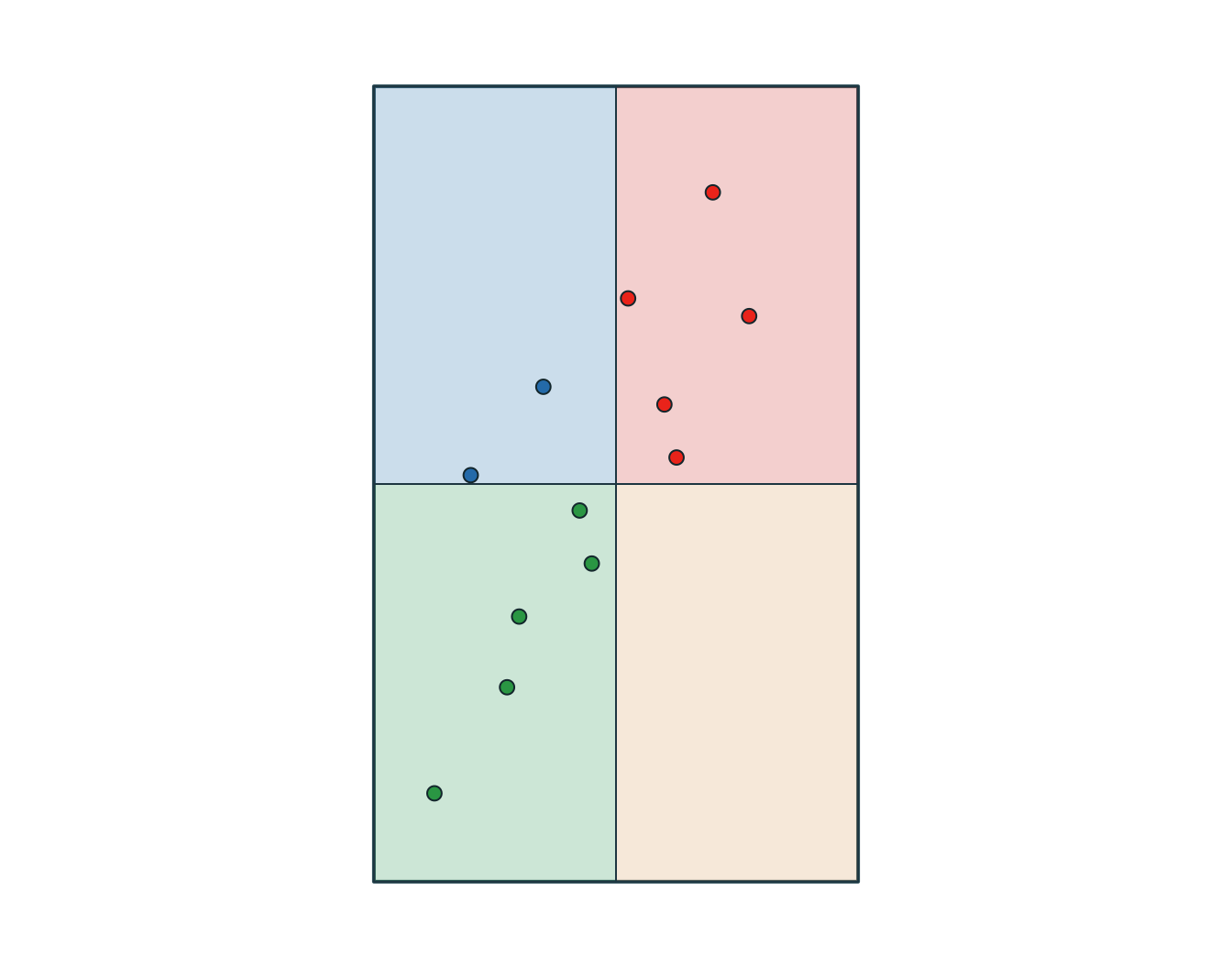
# A tibble: 6 × 3
track_id zone_id zone_type
<chr> <chr> <chr>
1 A NW forest
2 B NW forest
3 A NE urban
4 B NE urban
5 C NE urban
6 B SW wetland 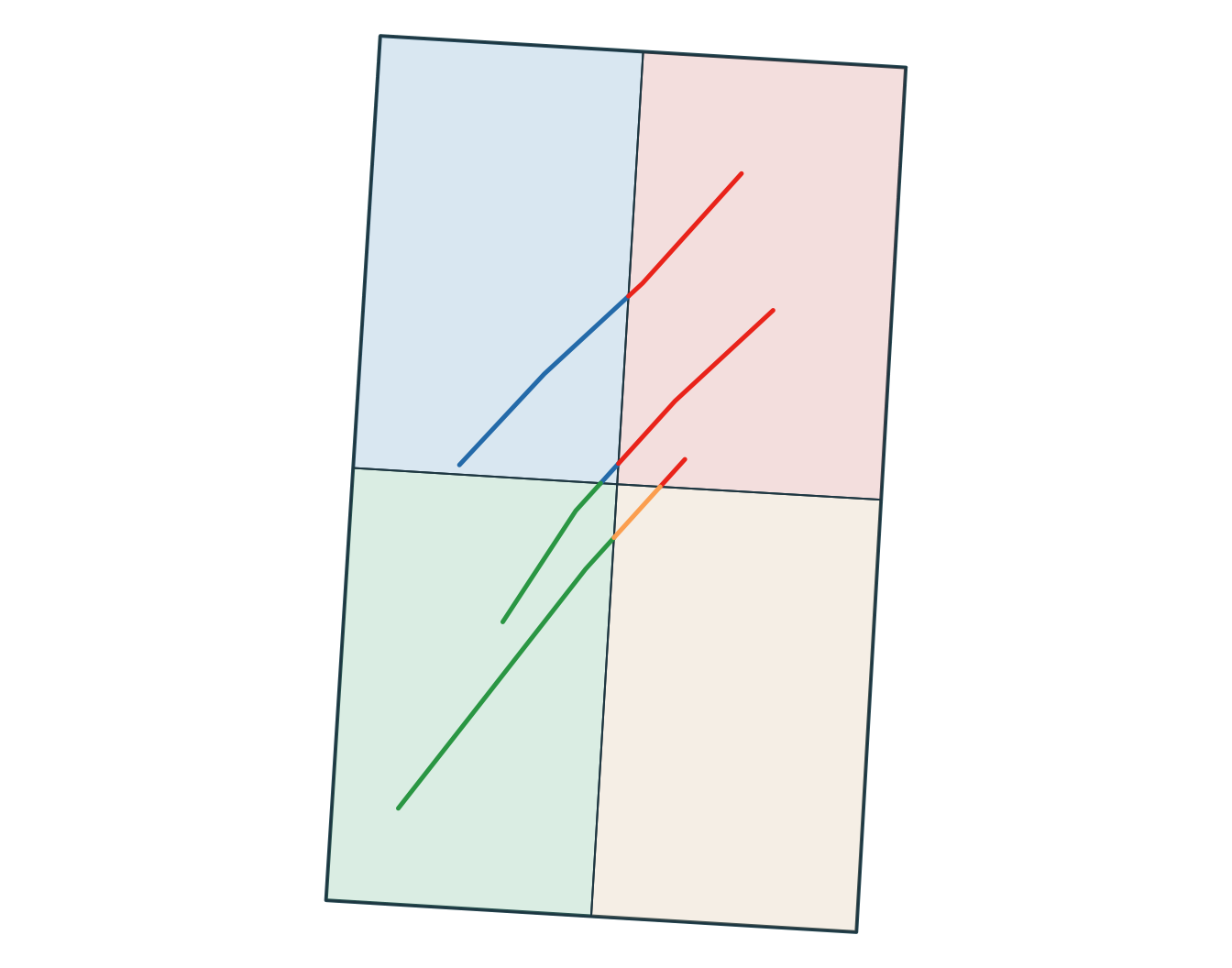
Note
Use a join when you want to attach attributes. Use an intersection when you want overlap to create new geometries.
[1] POLYGON
18 Levels: GEOMETRY POINT LINESTRING POLYGON MULTIPOINT ... TRIANGLE obs_id track_id seq timestamp lon lat category value
1 OBS01 A 1 2026-03-01 08:00:00 -72.92 46.78 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 -72.86 46.83 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 -72.79 46.88 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 -72.72 46.94 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 -72.88 46.70 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 -72.83 46.76 agriculture 13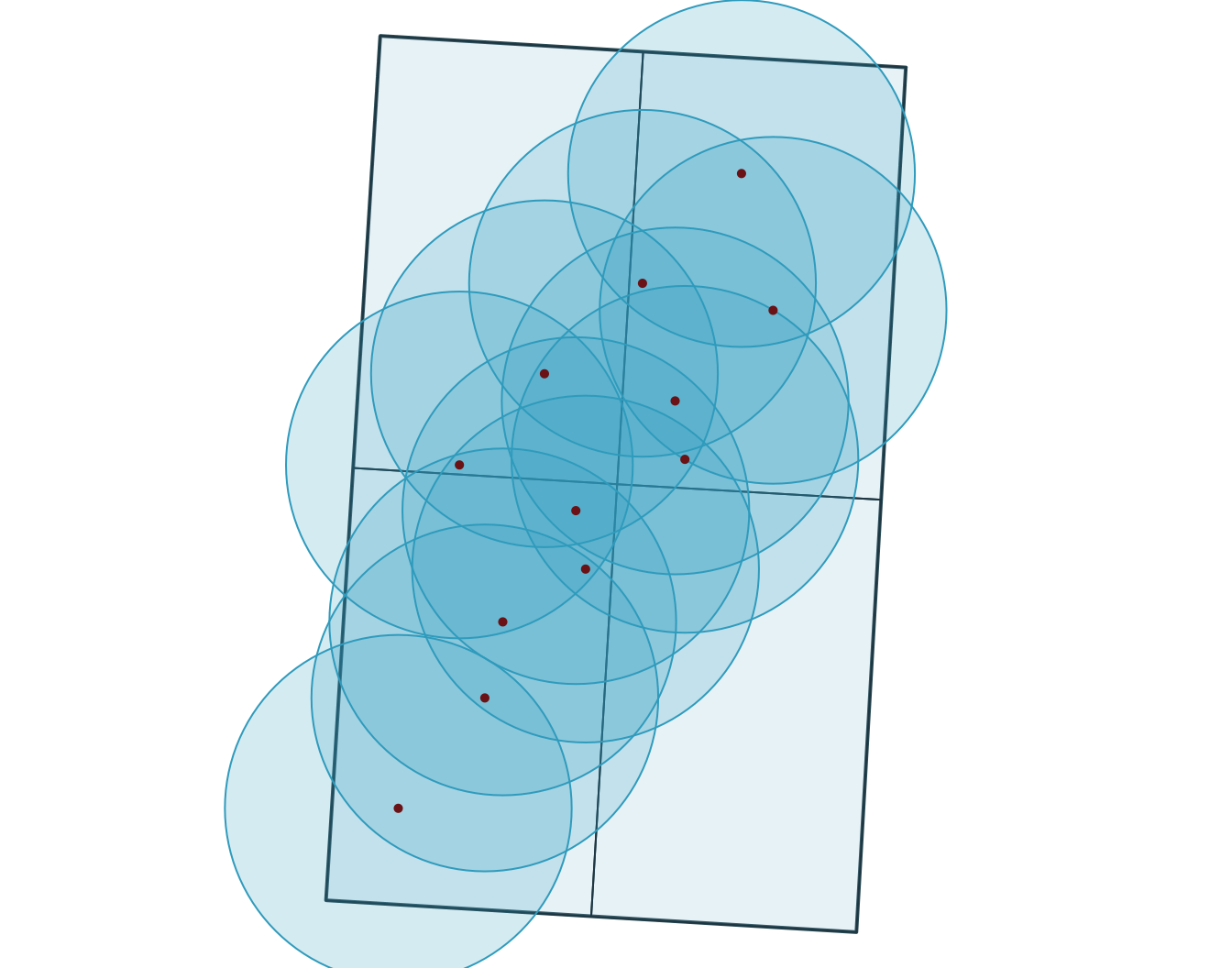
[1] "polygons" obs_id track_id seq timestamp category value
1 OBS01 A 1 2026-03-01 08:00:00 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 agriculture 13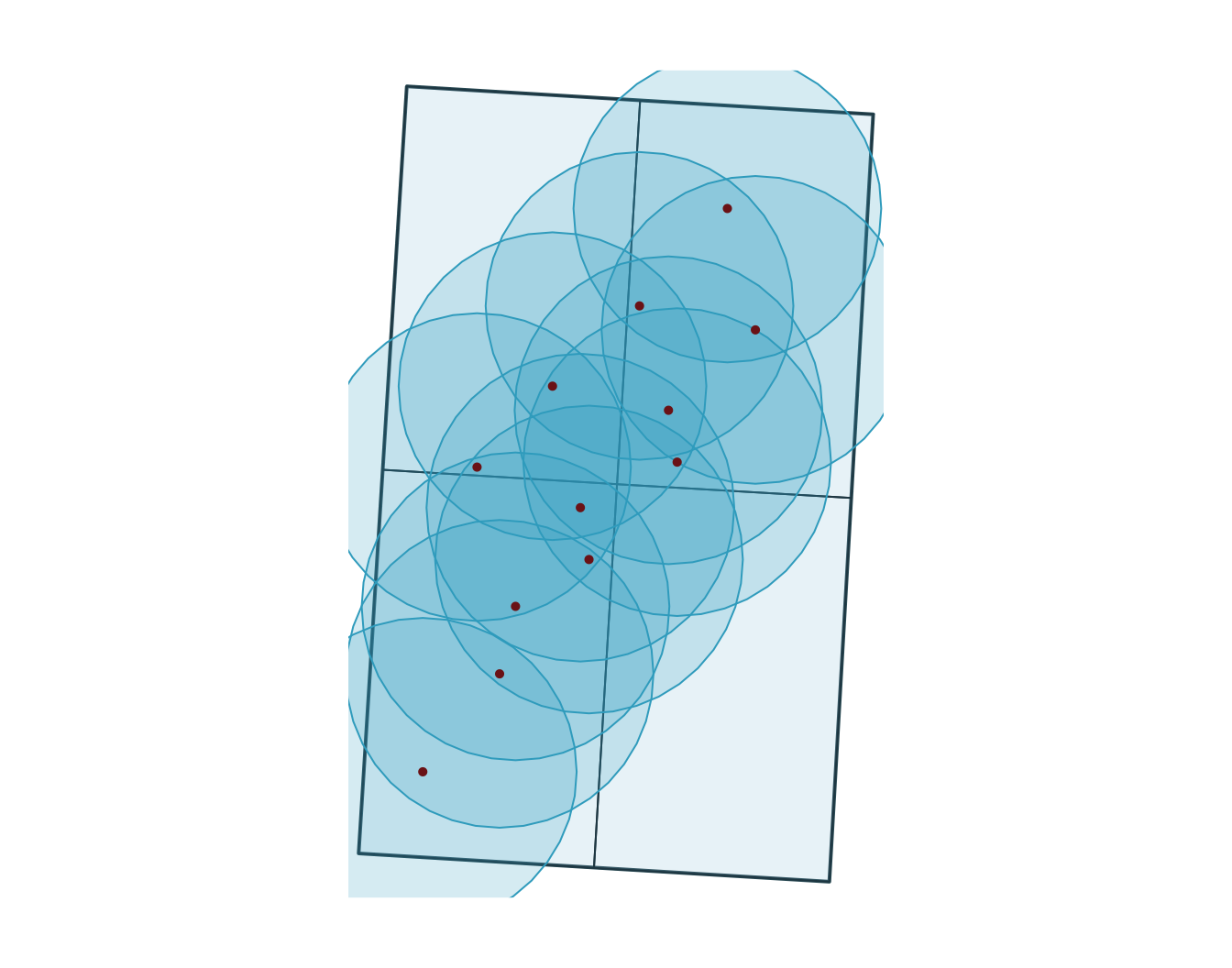
Note
Buffers create new geometries at a fixed distance around features, so they are usually done in a projected CRS with interpretable distance units.
Subset, measure, join, intersect, buffer
stars & terra
stars object with 2 dimensions and 1 attribute
attribute(s):
Min. 1st Qu. Median Mean 3rd Qu. Max.
surface.tif 9.6 40.8325 48.5 50.00007 59.805 100.32
dimension(s):
from to offset delta refsys point x/y
x 1 40 -73 0.01 WGS 84 FALSE [x]
y 1 40 47 -0.01125 WGS 84 FALSE [y]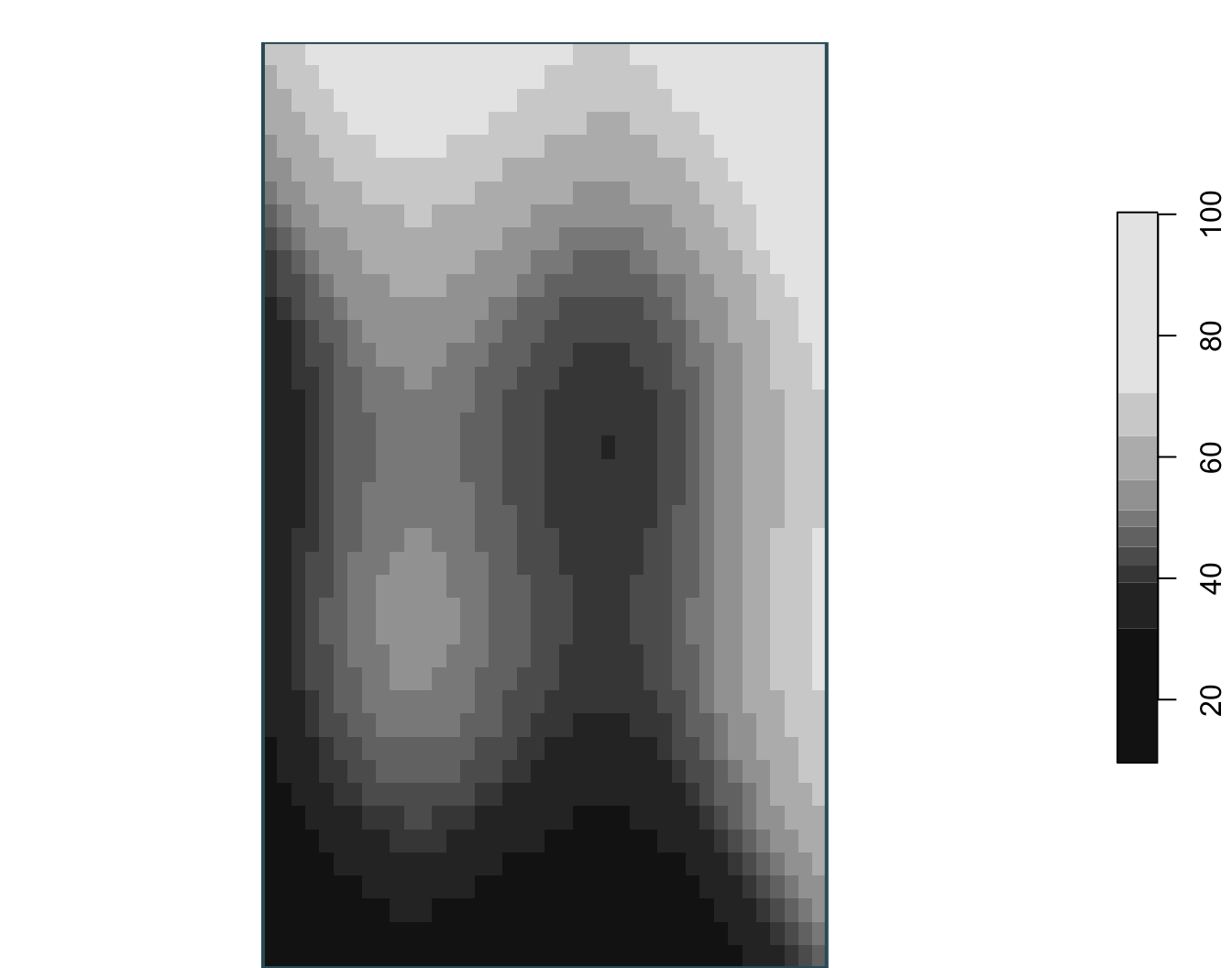
class : SpatRaster
size : 40, 40, 1 (nrow, ncol, nlyr)
resolution : 0.01, 0.01125 (x, y)
extent : -73, -72.6, 46.55, 47 (xmin, xmax, ymin, ymax)
coord. ref. : lon/lat WGS 84 (EPSG:4326)
source : surface.tif
name : surface_value
min value : 9.60
max value : 100.32 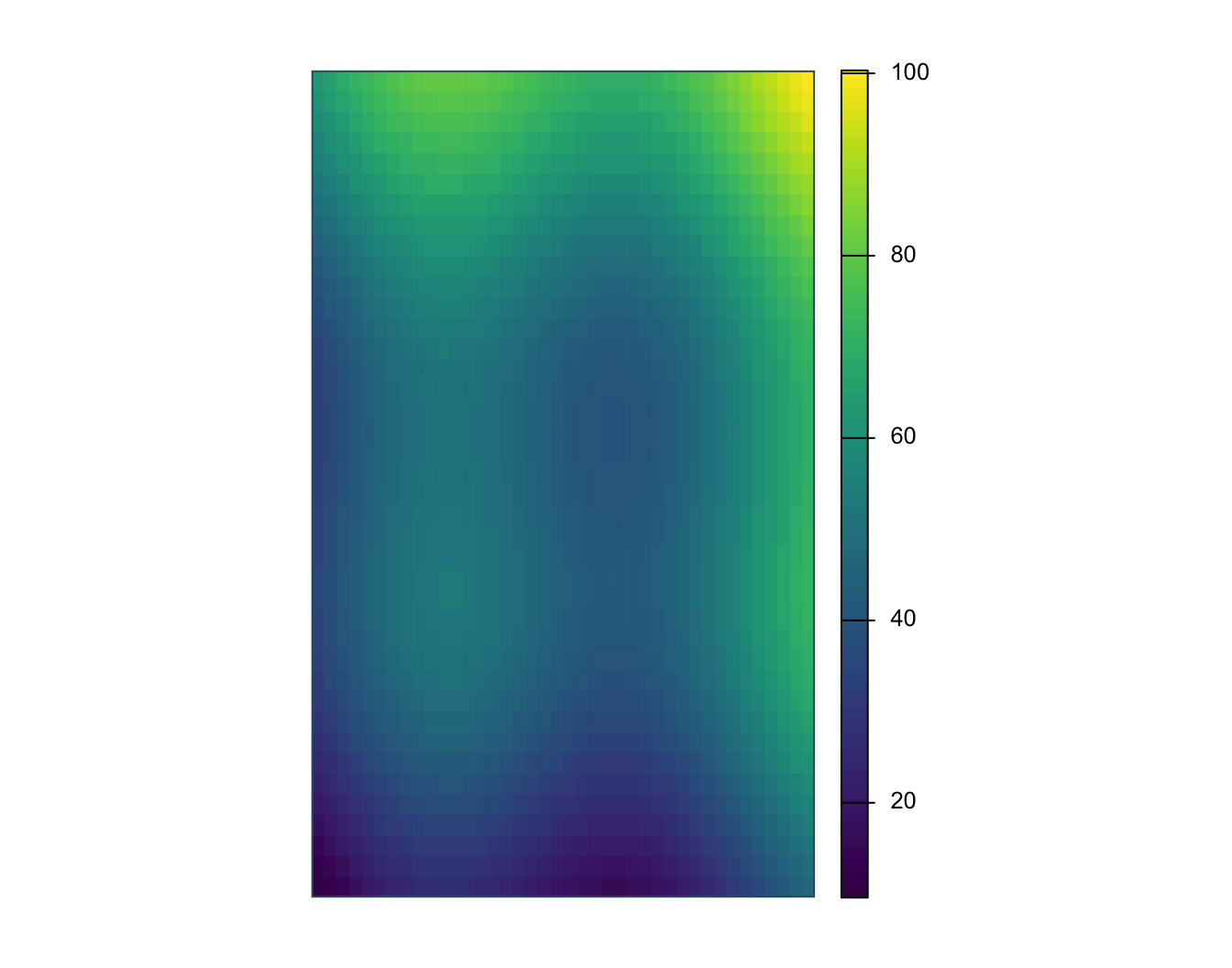
from to offset delta refsys point x/y
x 1 40 -73 0.01 WGS 84 FALSE [x]
y 1 40 47 -0.01125 WGS 84 FALSE [y] xmin ymin xmax ymax
-73.00 46.55 -72.60 47.00 [1] "WGS 84"Import, inspect, transform, export, visualize
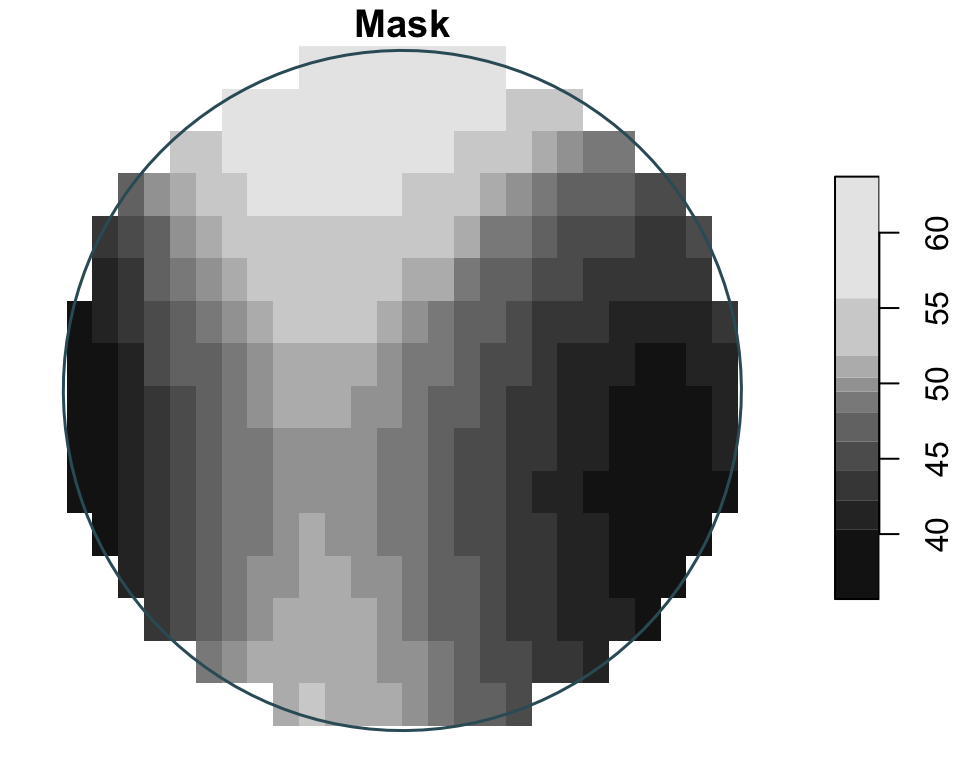
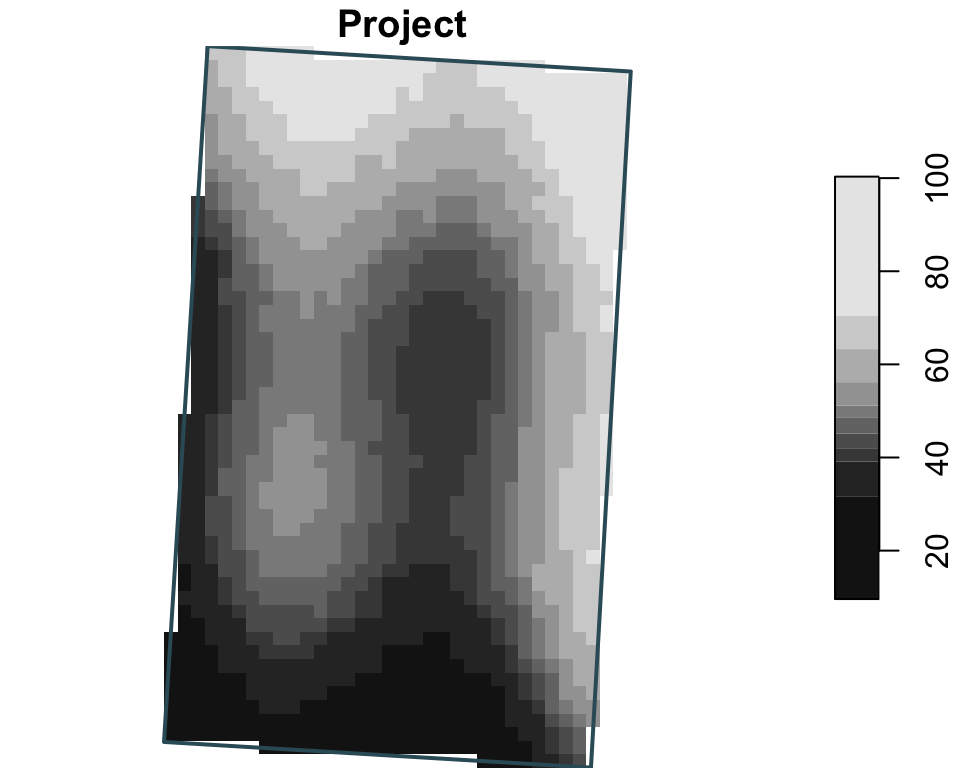

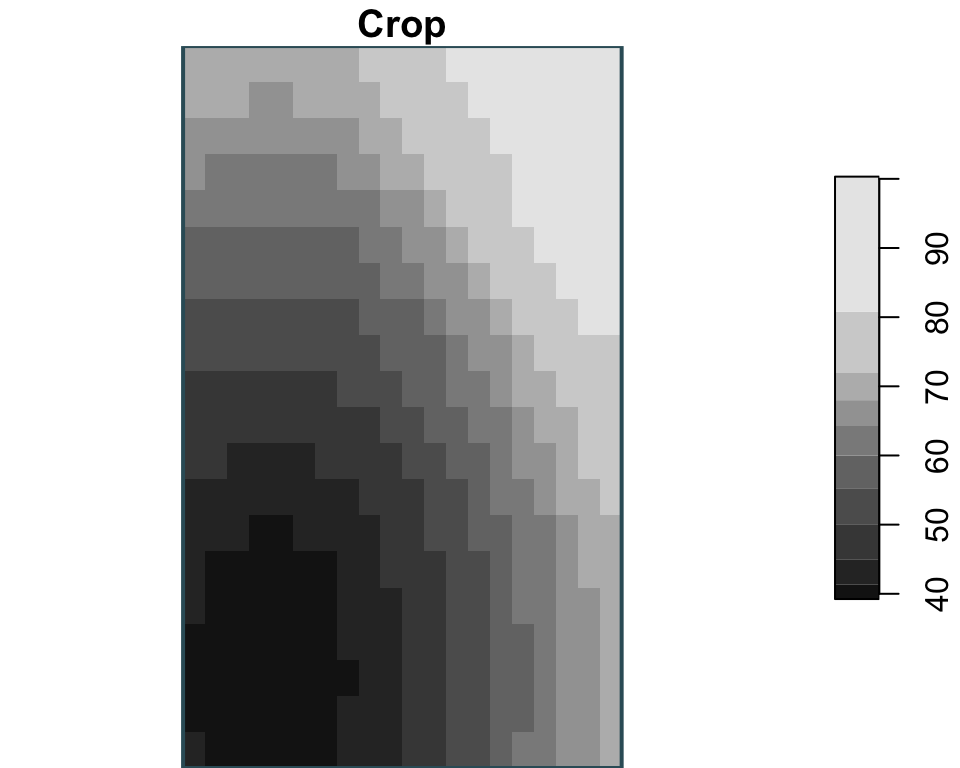

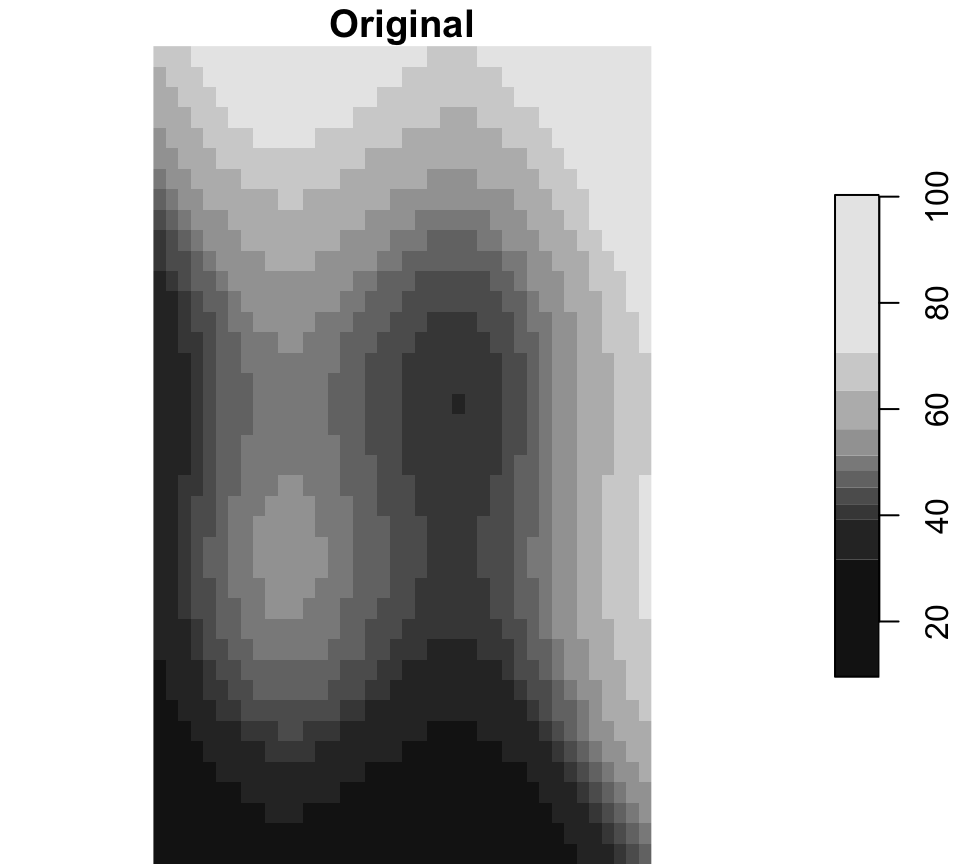
Raster manipulations
Here the three operations are shown independently: project changes CRS, crop uses one zone as the target extent, and mask keeps cells only inside a single buffer geometry.

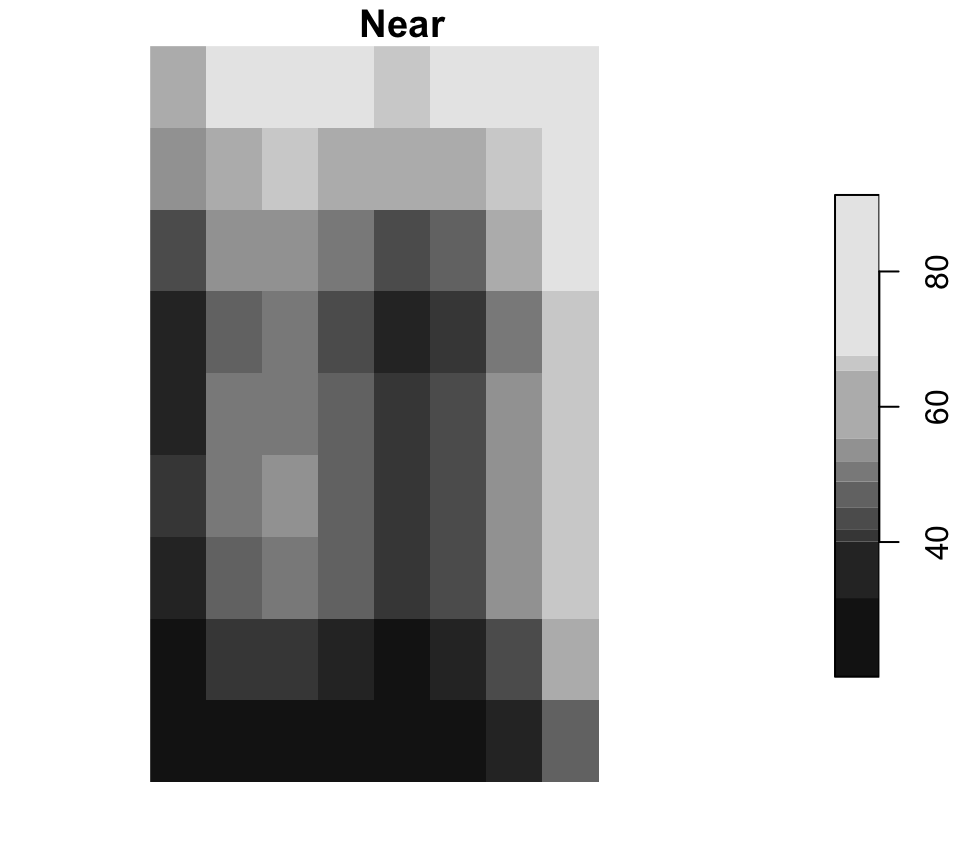
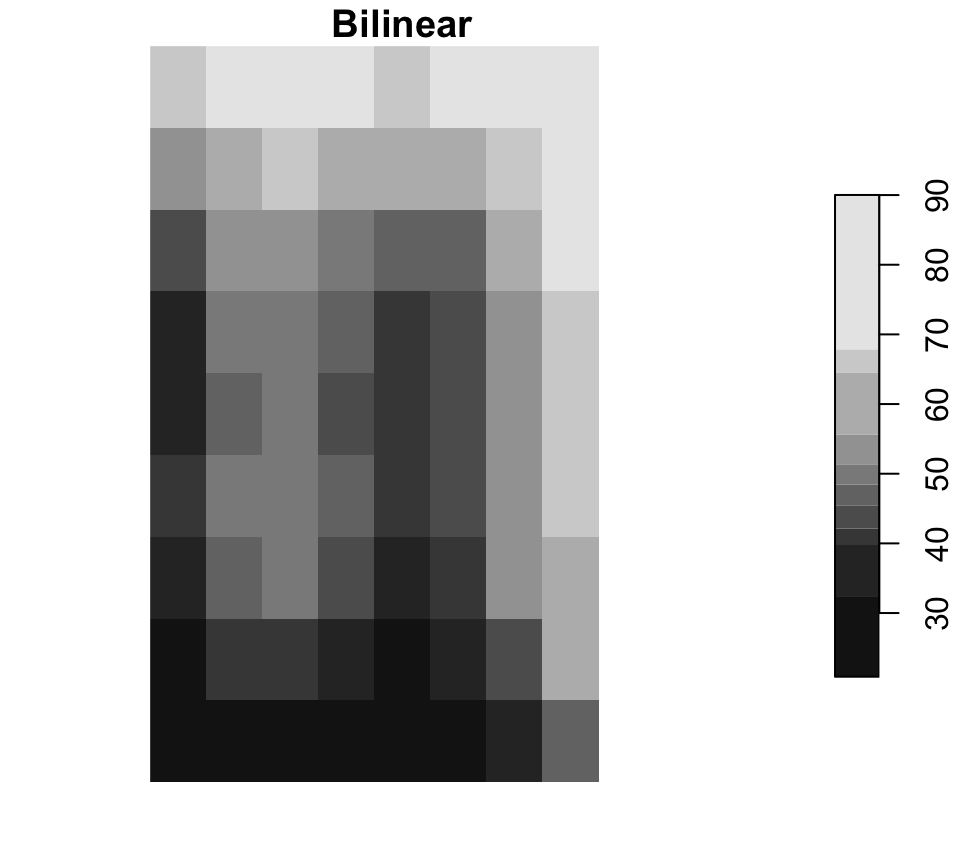
Resampling changes the cell grid
Near is common for categorical rasters, while bilinear is common for continuous surfaces.
[1] "surface.tif" "surface_alt"[1] 2
Multi-layer rasters
Can represent bands, variables, or time steps. Cell-wise functions combine information across layers.
Project, crop, mask, resample, layer operations
obs_id track_id seq timestamp lon lat category value
1 OBS01 A 1 2026-03-01 08:00:00 -72.92 46.78 urban 14
2 OBS02 A 2 2026-03-01 09:00:00 -72.86 46.83 urban 16
3 OBS03 A 3 2026-03-01 10:00:00 -72.79 46.88 forest 21
4 OBS04 A 4 2026-03-01 11:00:00 -72.72 46.94 forest 24
5 OBS05 B 1 2026-03-02 08:00:00 -72.88 46.70 agriculture 11
6 OBS06 B 2 2026-03-02 09:00:00 -72.83 46.76 agriculture 13
zone_id zone_type surface_value
1 NW forest 49.71
2 NW forest 48.70
3 NE urban 47.72
4 NE urban 61.39
5 SW wetland 51.67
6 SW wetland 45.88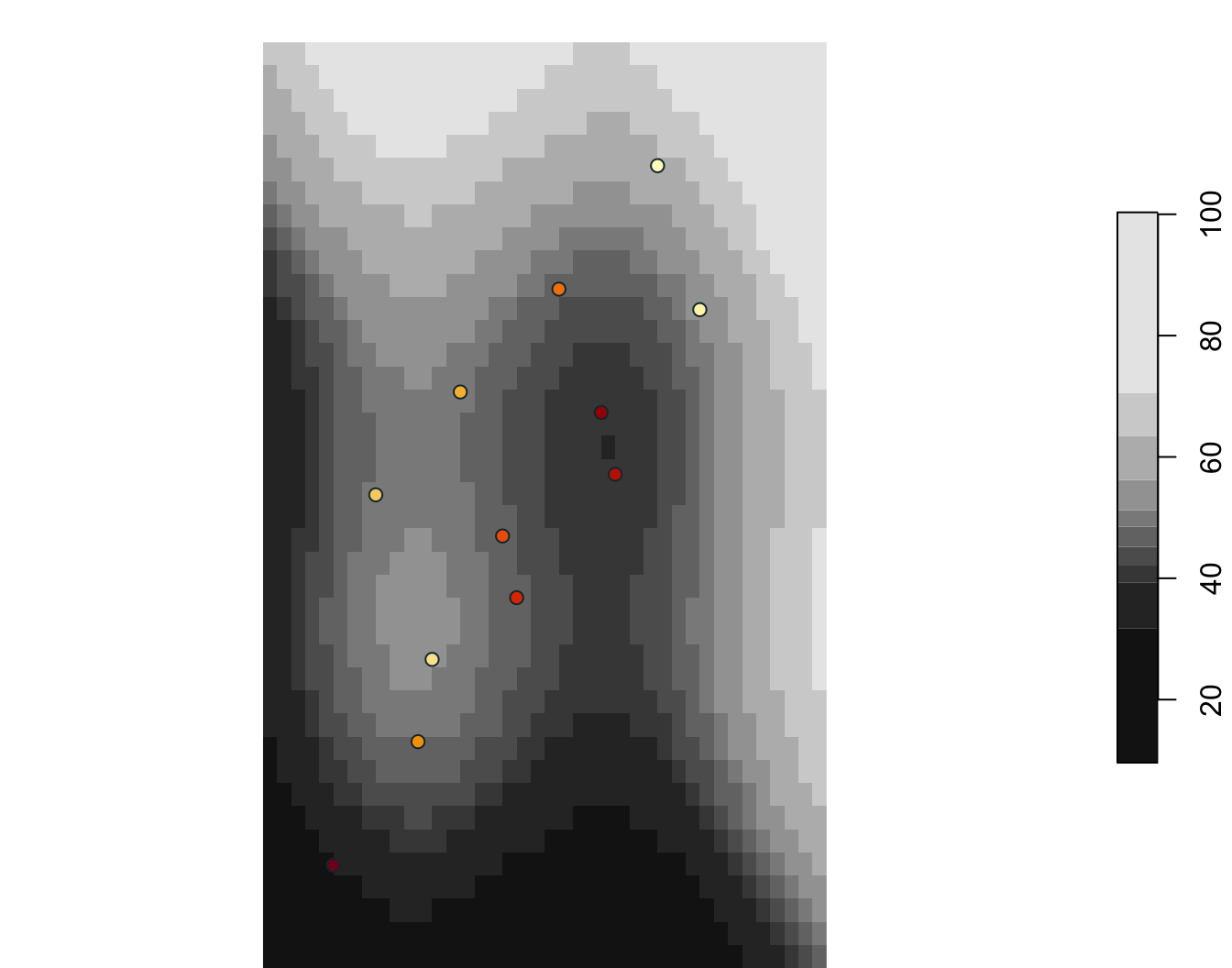
obs_id zone_id zone_type surface_value
1 OBS01 NW forest 48.69
2 OBS02 NW forest 49.60
3 OBS03 NE urban 47.72
4 OBS04 NE urban 59.98
5 OBS05 SW wetland 51.67
6 OBS06 SW wetland 47.16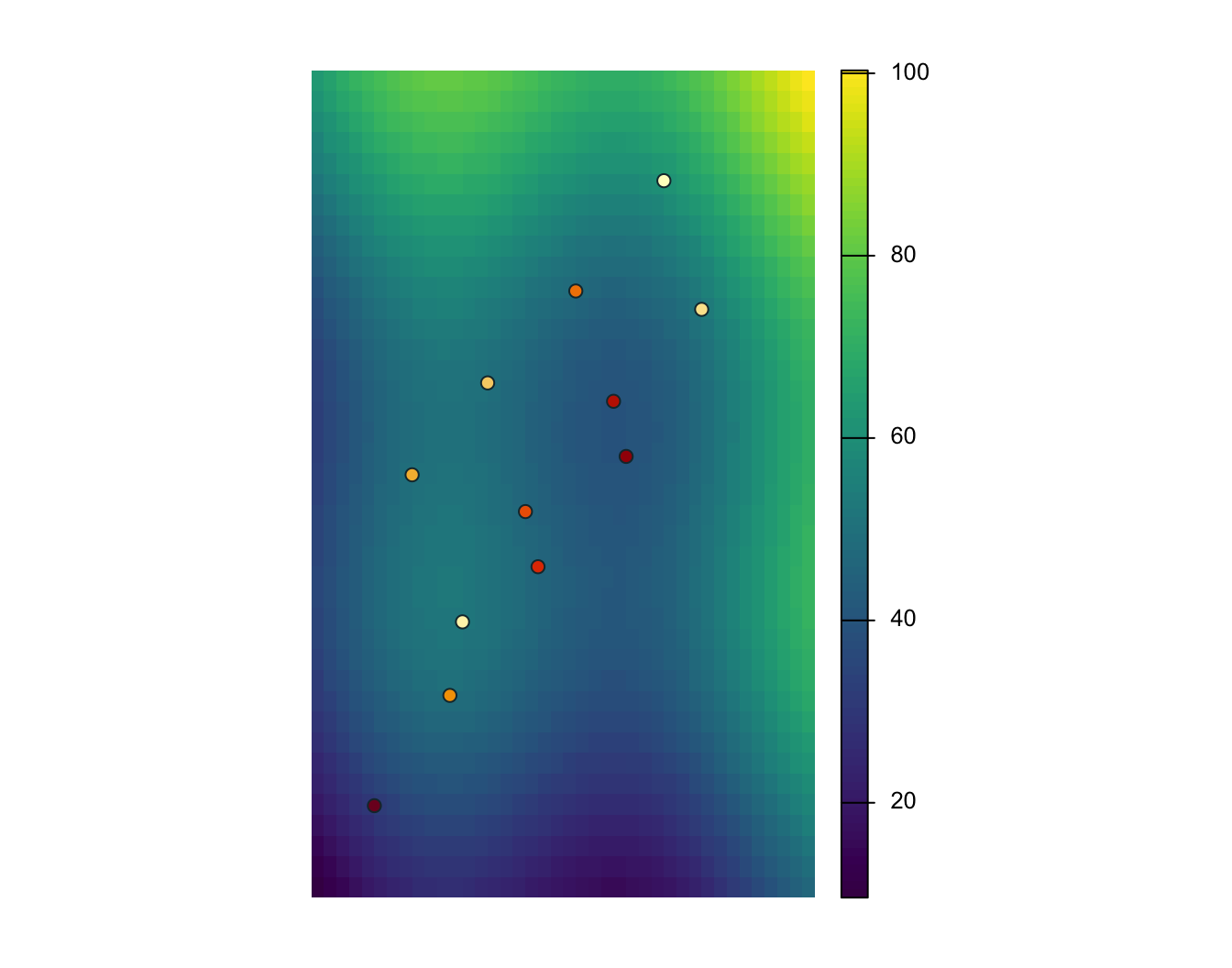
# A tibble: 3 × 2
track_id surface_value
<chr> <dbl>
1 A 50.0
2 B 45.0
3 C 44.2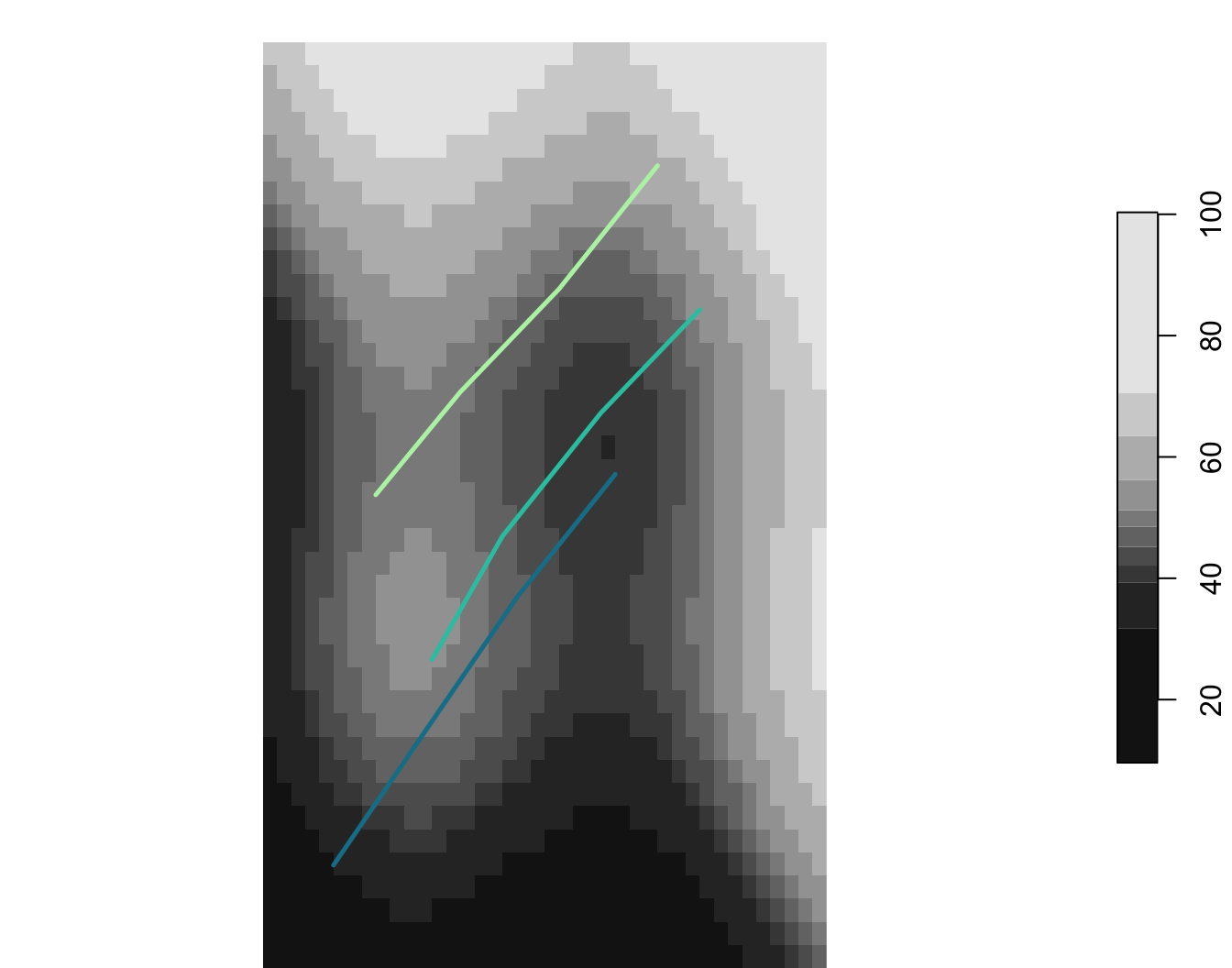
zone_id zone_type surface_value
1 NW forest 55.93852
2 NE urban 60.33677
3 SW wetland 39.66305
4 SE agriculture 44.06193
# A tibble: 3 × 5
zone_id zone_type point_n mean_surface mean_value
<chr> <chr> <int> <dbl> <dbl>
1 NE urban 5 48.4 19.8
2 NW forest 2 49.2 15
3 SW wetland 5 44.7 11.6# A tibble: 3 × 5
zone_id zone_type point_n mean_surface mean_value
<chr> <chr> <int> <dbl> <dbl>
1 NE urban 5 47.6 19.8
2 NW forest 2 49.1 15
3 SW wetland 5 44.5 11.6# A tibble: 3 × 6
zone_id zone_type point_n mean_surface mean_value area_km2
<chr> <chr> <int> <dbl> <dbl> <dbl>
1 NE urban 5 48.4 19.8 380
2 NW forest 2 49.2 15 380
3 SW wetland 5 44.7 11.6 382.# A tibble: 3 × 6
zone_id zone_type point_n mean_surface mean_value area_km2
<chr> <chr> <int> <dbl> <dbl> <dbl>
1 NE urban 5 47.6 19.8 381.
2 NW forest 2 49.1 15 381.
3 SW wetland 5 44.5 11.6 383.Extract, summarise, build analysis tables
The point here is not the analyses themselves. The point is the workflow that gets you from spatial data to analysis-ready inputs.
obs_id track_id zone_id zone_type category value surface_value
1 OBS01 A NW forest urban 14 49.71
2 OBS02 A NW forest urban 16 48.70
3 OBS03 A NE urban forest 21 47.72
4 OBS04 A NE urban forest 24 61.39
5 OBS05 B SW wetland agriculture 11 51.67
6 OBS06 B SW wetland agriculture 13 45.88 obs_id track_id zone_id zone_type category value surface_value
1 OBS01 A NW forest urban 14 48.69
2 OBS02 A NW forest urban 16 49.60
3 OBS03 A NE urban forest 21 47.72
4 OBS04 A NE urban forest 24 59.98
5 OBS05 B SW wetland agriculture 11 51.67
6 OBS06 B SW wetland agriculture 13 47.16 term Estimate Std..Error z.value Pr...z..
1 (Intercept) 2.178 0.536 4.064 0.000
2 surface_value 0.011 0.010 1.052 0.293
3 zone_typeurban 0.283 0.208 1.356 0.175
4 zone_typewetland -0.211 0.229 -0.921 0.357 term Estimate Std..Error z.value Pr...z..
1 (Intercept) 2.114 0.550 3.841 0.000
2 surface_value 0.012 0.011 1.144 0.253
3 zone_typeurban 0.292 0.209 1.399 0.162
4 zone_typewetland -0.205 0.229 -0.897 0.370pseudo_points <- st_sample(aoi_qc, size = nrow(points_qc), exact = TRUE) |>
st_as_sf() |>
st_join(zones_qc[, c("zone_id", "zone_type")])
pseudo_points$surface_value <- st_extract(surface_qc, pseudo_points)[[1]]
pseudo_points$presence <- 0
points_qc$presence <- 1
pa_data <- bind_rows(points_qc, pseudo_points) |>
st_drop_geometry() |>
filter(!is.na(zone_id), !is.na(surface_value))
glm_pa <- glm(presence ~ surface_value + zone_type, family = binomial(), data = pa_data) term Estimate Std..Error z.value Pr...z..
1 (Intercept) -16.407 3143.229 -0.005 0.996
2 surface_value -0.039 0.036 -1.098 0.272
3 zone_typeforest 19.471 3143.229 0.006 0.995
4 zone_typeurban 18.584 3143.229 0.006 0.995
5 zone_typewetland 18.917 3143.229 0.006 0.995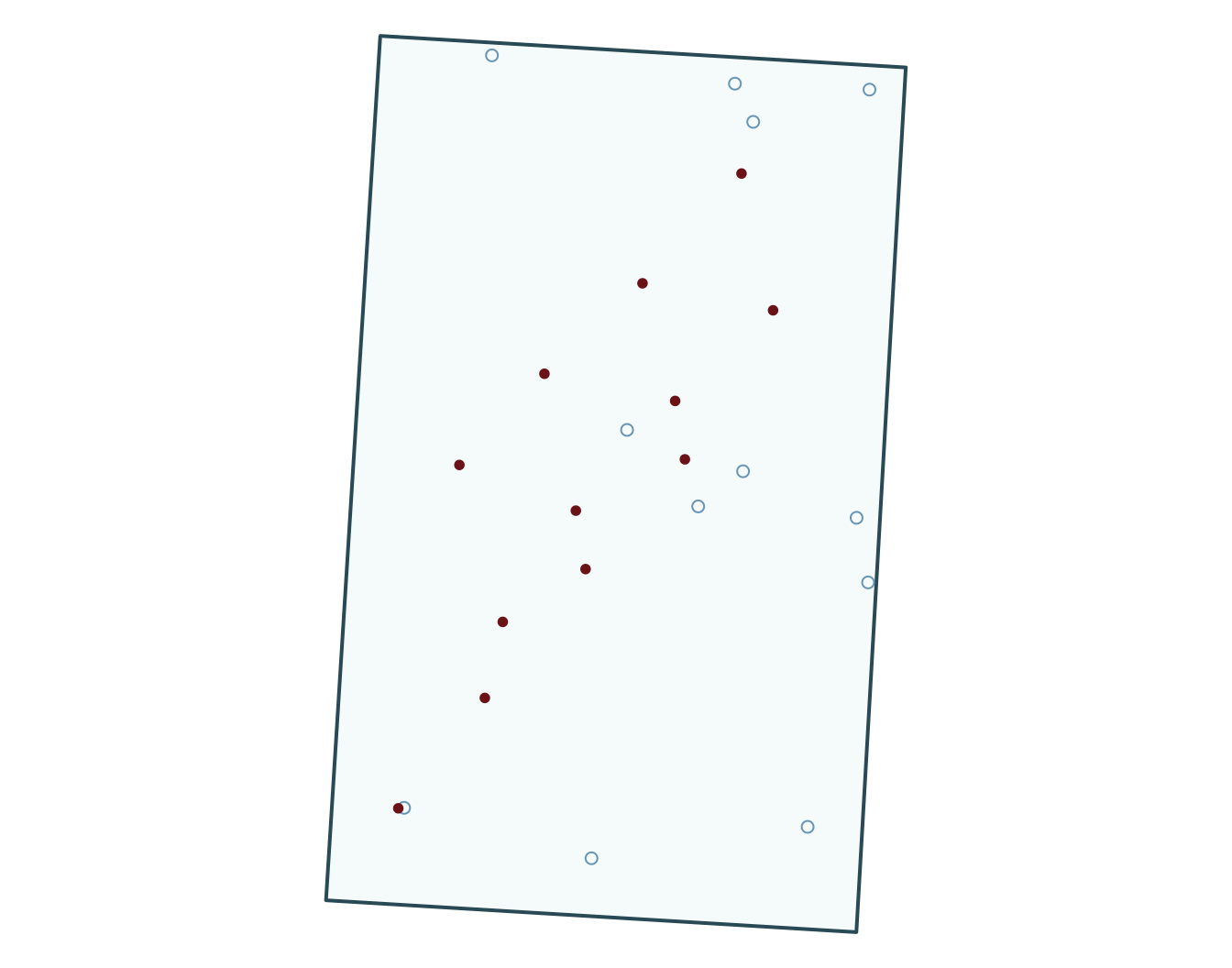
pseudo_points <- spatSample(aoi_qc, size = nrow(points_qc), method = "random", as.points = TRUE)
pseudo_zone <- extract(zones_qc, pseudo_points)
pseudo_surface <- extract(surface_qc, pseudo_points)
pseudo_points <- cbind(pseudo_points, pseudo_zone[, c("zone_id", "zone_type")], pseudo_surface[, "surface_value", drop = FALSE])
pseudo_points$presence <- 0
points_qc$presence <- 1
pa_data <- bind_rows(as.data.frame(points_qc), as.data.frame(pseudo_points)) |>
filter(!is.na(zone_id), !is.na(surface_value))
glm_pa <- glm(presence ~ surface_value + zone_type, family = binomial(), data = pa_data) term Estimate Std..Error z.value Pr...z..
1 (Intercept) -13.913 2782.290 -0.005 0.996
2 surface_value -0.061 0.062 -0.981 0.326
3 zone_typeforest 16.642 2782.287 0.006 0.995
4 zone_typeurban 17.569 2782.287 0.006 0.995
5 zone_typewetland 17.009 2782.287 0.006 0.995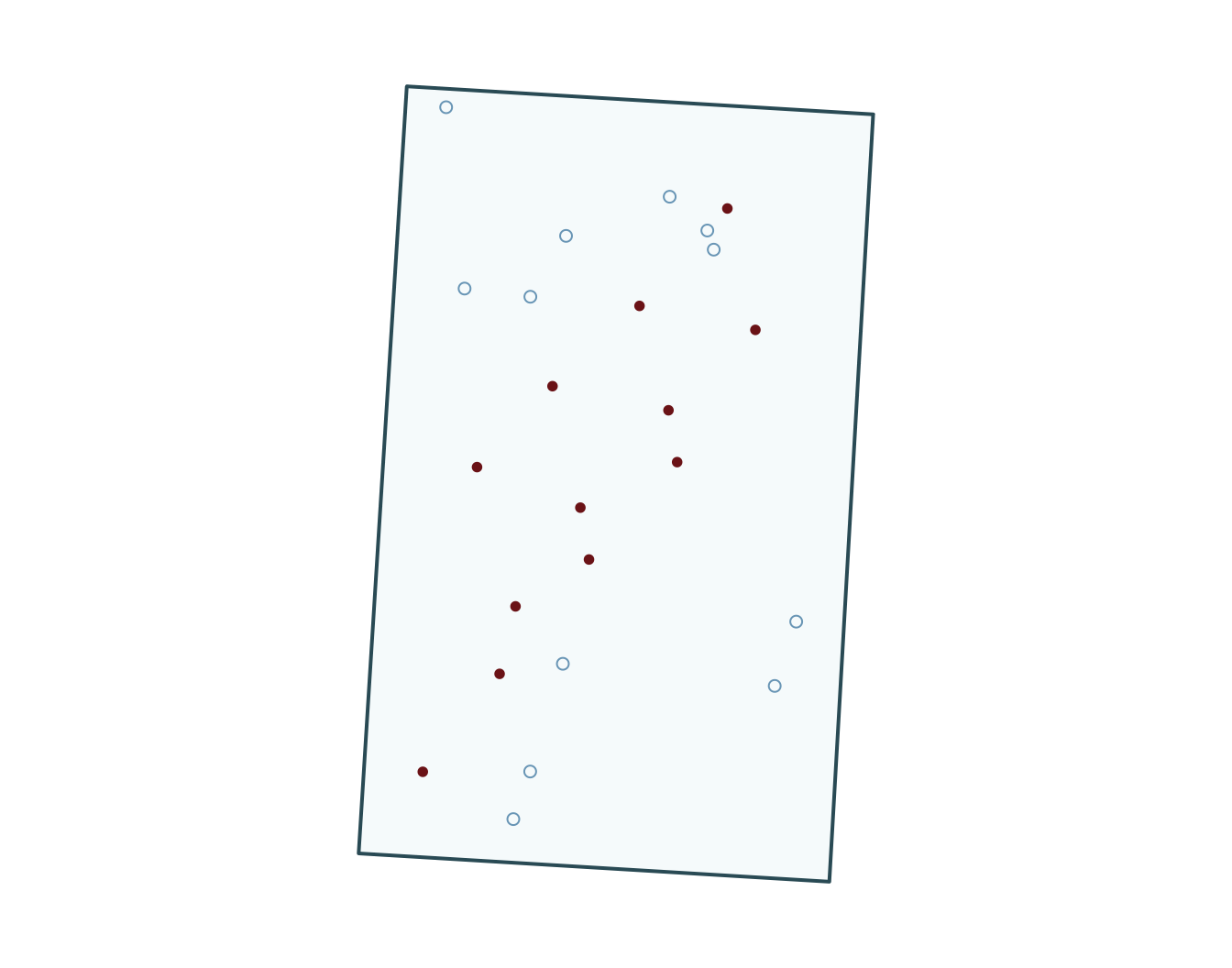
Prepare data, fit simple GLMs
Bonus: generate pseudo-absences for your GLM with points data
Use as an exploratory tools
$x_head
[1] -344515.3 -344091.9 -343668.5 -343245.1 -342821.7 -342398.3
$y_head
[1] 292854.5 293508.8 294163.2 294817.5 295471.8 296126.1
$z_dim
[1] 80 80$x_head
[1] -344515.3 -344091.9 -343668.5 -343245.1 -342821.7 -342398.3
$y_head
[1] 292854.5 293508.8 294163.2 294817.5 295471.8 296126.1
$z_dim
[1] 80 80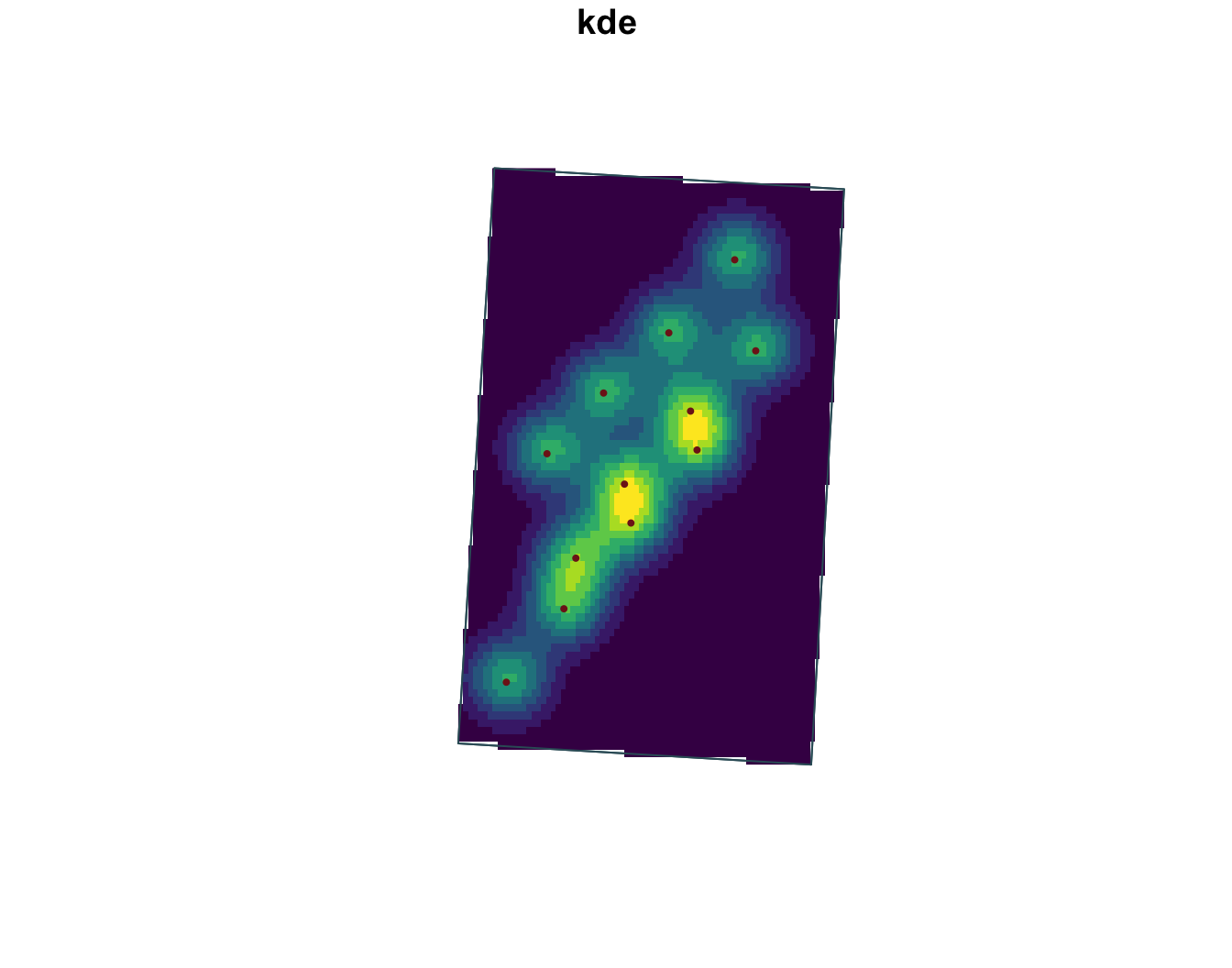
Fit KDEs and explore surfaces
Quick exploration
plot()mapview()Communication
ggplot2tmapUse exploratory maps to inspect data quickly, then move to more deliberate mapping tools when the goal is communication or sharing.
ggplot2ggplot2 works especially well for polished static maps, particularly with sf and stars objects.
gg <- ggplot() +
geom_stars(
data = points_kde_sf_plot
) +
scale_fill_viridis_c(name = "KDE") +
geom_sf(
data = zones_qc,
fill = NA,
color = "#355c67"
) +
geom_sf(
data = aoi_qc,
fill = NA,
color = "#17313b"
) +
geom_sf(
data = points_sf_qc,
aes(color = category),
size = 1.7
) +
coord_sf(expand = FALSE) +
theme_minimal()
gg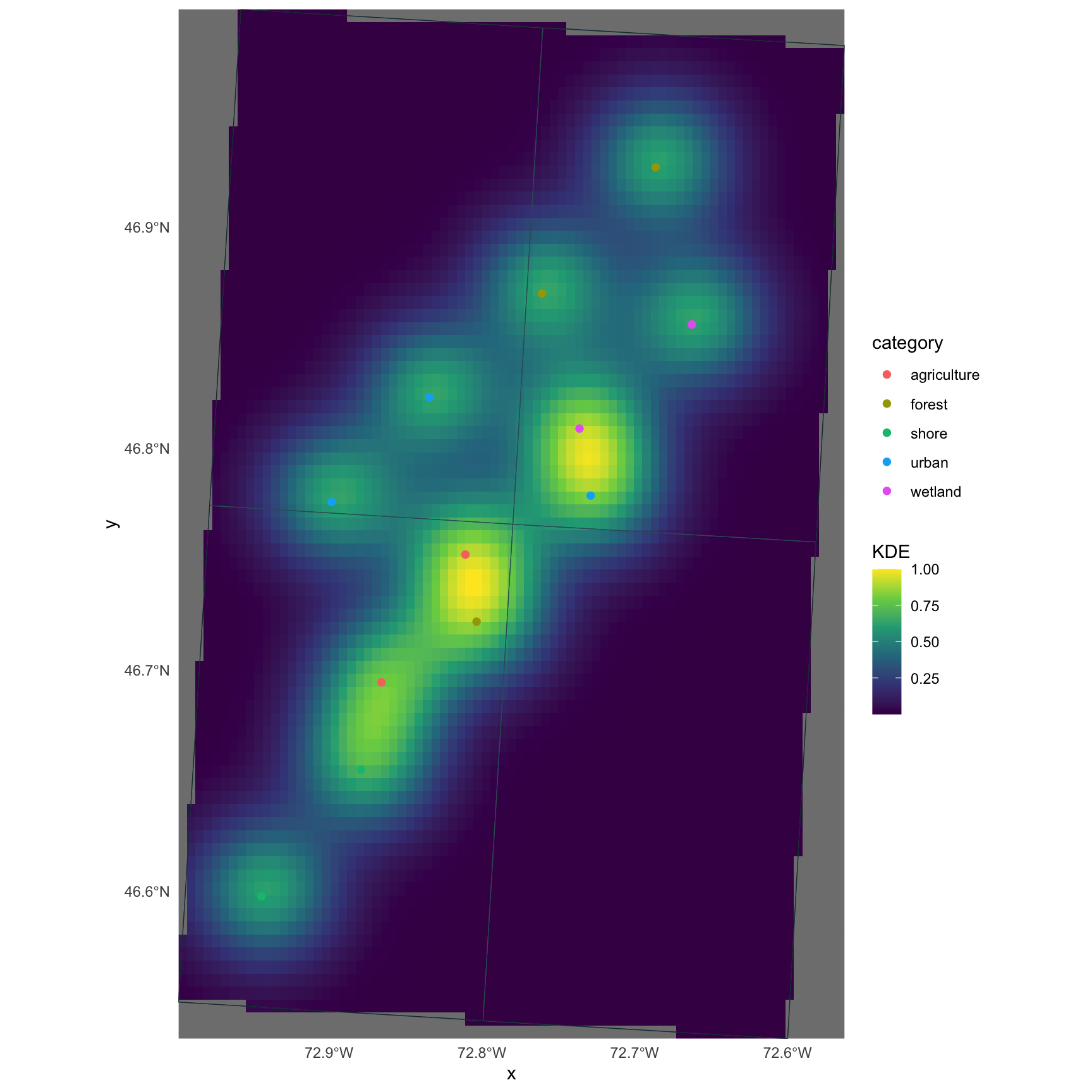
tmaptmap is built for thematic mapping and works well when you want a more map-focused grammar than ggplot2.
tm <- tm_shape(points_kde_sf_plot) +
tm_raster(col.scale = tm_scale_continuous(
values = "viridis"
)) +
tm_shape(zones_qc) +
tm_borders(col = "#355c67") +
tm_shape(points_sf_qc) +
tm_symbols(
fill = "category",
size = 0.5
) +
tm_shape(aoi_qc) +
tm_borders(col = "#17313b") +
tm_layout(
frame = FALSE,
legend.outside = TRUE
)
tm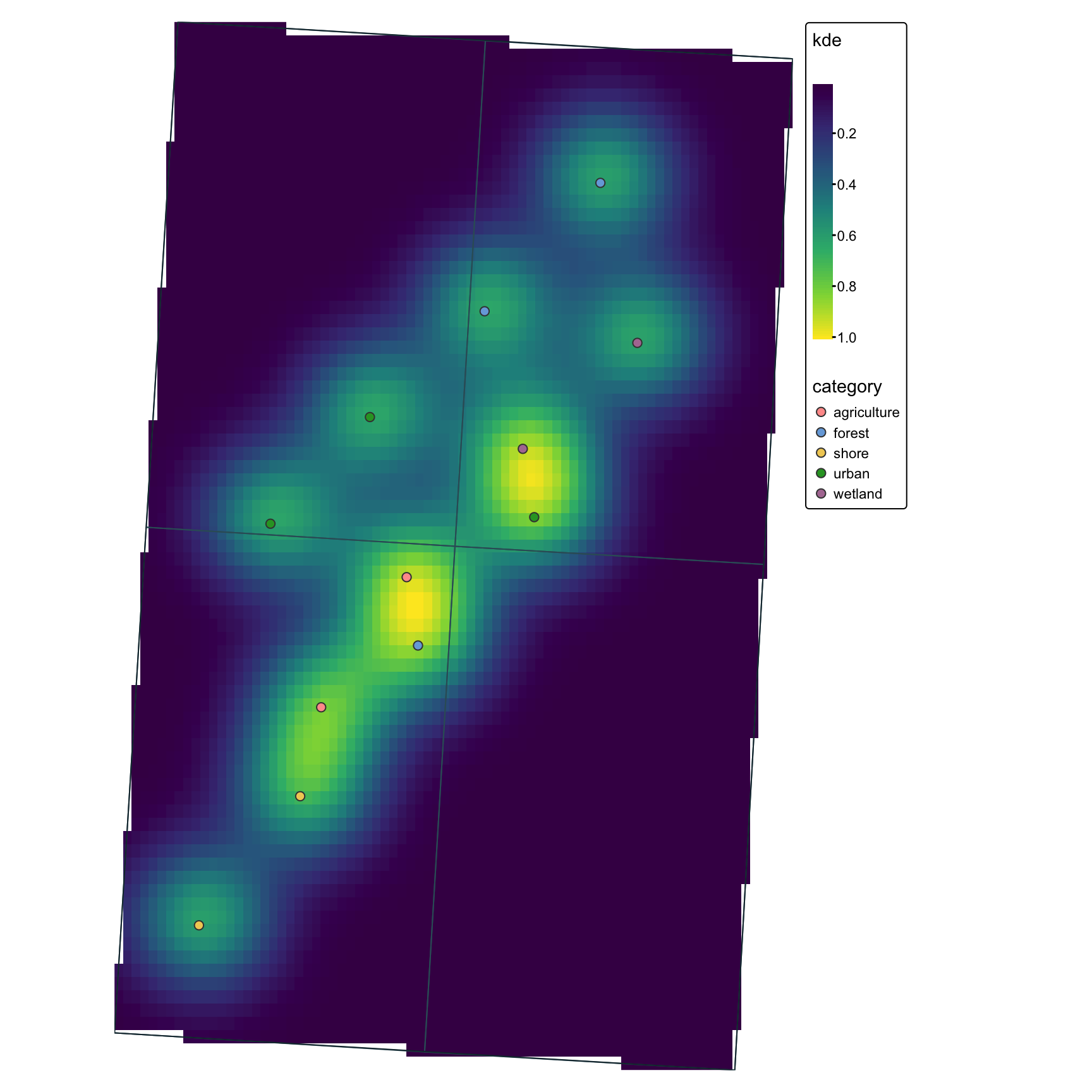
tmapOne advantage of tmap is that the same map logic can often be used for static output or for an interactive shared map.
output file tool
1 presentation figure advanced_map.png ggsave(..., dpi = 300)
2 publication-ready figure advanced_map.pdf ggsave(...) output file tool
1 static thematic map advanced_map.png tmap_save(...)
2 interactive shared map advanced_map.html tmap_mode("view") + tmap_save(...)Build, export, and share maps
sf, stars, and terra cover most common spatial workflows in Rprompts: